Large Scale Sequence Analysis with Applications to Genomics
|
|
|
- Mari Kapulainen
- 6 vuotta sitten
- Katselukertoja:
Transkriptio
1 Large Scale Sequence Analysis with Applications to Genomics Gunnar Rätsch, Max Planck Society Tübingen, Germany Talk at CSML, University College London March 18, 2009 Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
2 Discovery of the Nuclein (Friedrich Miescher, 1869) Tübingen, around 1869 Discovery of Nuclein: from lymphocyte & salmon multi-basic acid ( 4) If one... wants to assume that a single substance... is the specific cause of fertilization, then one should undoubtedly first and foremost consider nuclein (Miescher, 1874) Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
3 of the DNA of C. elegans >CHROMOSOME I GCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGC CTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCT AAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAA GCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGC CTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCT AAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAA GCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGC CTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCT AAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAAAAATTGAGATAAGAAAA CATTTTACTTTTTCAAAATTGTTTTCATGCTAAATTCAAAACGTTTTTTT TTTAGTGAAGCTTCTAGATATTTGGCGGGTACCTCTAATTTTGCCTGCCT GCCAACCTATATGCTCCTGTGTTTAGGCCTAATACTAAGCCTAAGCCTAA GCCTAATACTAAGCCTAAGCCTAAGACTAAGCCTAATACTAAGCCTAAGC CTAAGACTAAGCCTAAGACTAAGCCTAAGACTAAGCCTAATACTAAGCCT... Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
4 Research Topics Machine Learning 1 Inference methods for structured data Develop fast and accurate learning methods 2 Convergence properties of iterative algorithms Boosting-like algorithms and semi-infinite LPs 3 Genome annotation Predict features encoded on DNA Molecular Biology 4 Biological networks Understand interactions between gene products 5 Analysis of polymorphisms Discover polymorphisms and associate with phenotypes Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
5 Research Topics Machine Learning 1 Inference methods for structured data Develop fast and accurate learning methods 2 Convergence properties of iterative algorithms Boosting-like algorithms and semi-infinite LPs 3 Genome annotation Predict features encoded on DNA Molecular Biology 4 Biological networks Understand interactions between gene products 5 Analysis of polymorphisms Discover polymorphisms and associate with phenotypes Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
6 Inference Methods for Structured Data 1 Large scale sequence classification with Sonnenburg (Fraunhofer, Berlin) & Schölkopf (MPI Biol. Cybernetics) 2 Analysis and explanation of learning results with Sonnenburg (Fraunhofer, Berlin) 3 Sequence segmentation & structure prediction with Altun (MPI Biol. Cybernetics) Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
7 Inference Methods for Structured Data 1 Large scale sequence classification with Sonnenburg (Fraunhofer, Berlin) & Schölkopf (MPI Biol. Cybernetics) 2 Analysis and explanation of learning results with Sonnenburg (Fraunhofer, Berlin) 3 Sequence segmentation & structure prediction with Altun (MPI Biol. Cybernetics) k mer Length Position Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
8 Inference Methods for Structured Data 1 Large scale sequence classification with Sonnenburg (Fraunhofer, Berlin) & Schölkopf (MPI Biol. Cybernetics) 2 Analysis and explanation of learning results with Sonnenburg (Fraunhofer, Berlin) 3 Sequence segmentation & structure prediction with Altun (MPI Biol. Cybernetics) k mer Length Position Log-intensity transcript Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
9 Computational Genome Annotation Simplest formulation: Given a DNA sequence x { A, C, G, T } L Find the correct label sequence y = y 1 y 2... y L (y i Y = { intergenic, 5 UTR, coding, intron,... }) Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
10 Example: C. elegans (I: 43,500-52,050) GAAGAAATGGAGCATTTGCGCTCCATCACACTCTCAGACAATTTCATTTTCCACATCCTATATATATTTTGGTTTTTCTGTCGTATTTTGTTTTAATTTATTGGTATTTCGTTCAAAAATAATTATTTTGACTGTATTTTTGGTTGCATA CATGTAGAACTGCTGTTTTTTAAGATATTCTGCCCATTCAAGTTTTTCAGTGTAAAATTGATATATTTCATTCCAACTGAAAATGAGATCGAAACGATGGAAAACCTCGGATATTACTGATTATGGAAAGAAGAGAAAAGAATCGGAAAG TTGTGGATCAAGTTCACCGATTCTCGAAACACAGTCATCTGGCGGTGCGGAACTTGACGAAGTTACTGAGGATGAATATTCTAGTAATTCGAGCAGTAATGAAACTAGCGACGAAGAGGAAAACTCAGAAGTACCAAATGTCTTATCTAT AACAGAAAGAGGTAAGAATTGCGTCTTCTAGTGATCATACTTTTCGCCAGATTCCCTAATGTAATATATTTTGTTGTAGAGAAAAGTTGGCAAAAGTTAACGGAAAACGATTTGGGACGAATTCGTTTCATCTTGAAGTACACTAGCAAT ACTAAAAAATGCGTGAACGAGTATTTTCAATATAATCATGGGCAAAACAATGAAATTATGAAAAGTCTATTATTGGATACCGATGGAACTATGACTGCAAAGGCTTGTTCGGAATGTGCCTACGATTTGAATCAGTAAGTTACTCTCTCG ATTTATTCCCAAAATTAATATGTGCTTCAGGTGCCACTGCAAAAAACCGCTTCGCTTCATCAATGCTCCGTGTGGTTGGTTTGCTATTCAAAACTATAAATAGTTCACTGTTTCCGTTCAGAGGTCATCAACCAAGTTCTTCATGTTGAA AATGCGGAGCCCACCAGGATCAACCATGTAATCGCAACACTCTTCCGGAATCACATTGGCGAGATTTTGTTGGTCCACTCTATTTCTGTGCGAGAACTGTGATAAAACTAGTATTTTCAGCACAAAGGCTCGAACTGCGGAAGCTCGCGC ATCTGAAGAAGCTCAAATCAGGATTCAAATCCAAGACAACTCGAACGCATTCCAAAGATCGTATCATAACGATCCACAACCTTCATCAGCCGAAGAACATGAGGAAGATATCGTGGTGGATGGCTGAGTACGGAGCTCAAATGCCTTAAG GCGAAACAATTGGTTTTTTAATTTGCTGGTTATCATGTTAGATTTTGAACGTGTTAGGTCTTTCAATTGTTTTTTTTTTTCGAAATGTTGTTGTTCTAATAAATTTGTTTTATTTAATCAAACGTTTTTTAGTCTACTACGGGCGTGAAG CCAGATATCAGTGGTATCTTCTTATCAGAAGCTGAATCATTTCCGGTTGACAATGTTTGAAGGACATAAGAAAGGCTGTGTTACTGATTTCGACCATTGATTTGTTTATATATGGATATGTTCCACTGCCTTTTGGAAAGGCAGTATTCC CGGTATATATGGGCCTAATACGGAATCTAAAATAACCTGACACAAACCTGACGTTGACCTGTTGCCGGCCCGCGGCGGCTTAGTGTCAACTTGACAGCGGGTCGCGATTTCACCTGCCAGTTGTTCTCCATTCAGCAGCCAGCGACCTGC TGGCAGGTTGCCACTAACCTGACGCGGTTTACCTGTGTTATCGGCGCGTGCATAGCTTAGTGGTTTCAGGAAATGATGCTAGTAATCAGAAGATCGGGGTTCGGGAAACGGCAGGGGCTTGAAGGTTAGGTTCTATGAAGCAGGGCGAAG GGTTGACAAGGAGAGGCAATAAGCAAGTAGTAGGGGTTCTCTAGAAAACATTTTTGTCTTTAATATGCGTTTCCTACTGATTTATTATTGATATTTGGATCCCCTTTTCTAGAAAAAAAAATCAGAATCAGCAGAAAAATTTGAGAAAAA GTCATAGCAAATCAGAGTTGGTCAGAGTAAATCAGAGCTAGTCATAGTAAATCATAGCTAGTCAGAGAATATCAGAGTTAATCAGGGTAATAAGTAGACCTAGTCATAGTAAATCAGAGCTAGGCATAGTAAAGCGTGGTTACTCCGAGT AAAACCACACTTGCACCGAACTGCGGTTAGTGTGCTTTACCATTATGTAACTCCGCTTTTTACTCTGAGTTAGTATGATATGGTTTGTCTGAGCTGTGGTTGGGCTTCGCGGGAAACTTGAATAATTCGAGACAAAATCTAATTTTAGCG AATTTTCTTTAATTTCTTTGAGGTTTCTACGACAGAACTCGAAAAATTTCGGGTTTTAATGTTTACACATTTTATTTAAAATTGAATAATCAACTGCGGGACTCCTCGAAAATCACATGCTCATTTAAATTTTGAAGTTCAAACCTCAAA AAACGCGCAAAAACCAAATTCAGCTAGGATATCAAATTTATGATTGAAATCTATATTTTGATGCGGTGTTTCTGAAGTTTTCGCGATAAAATCCGAATAATAATTCCACGTACCGTATATTCTCTATCTAATTTCCAGGTCATTTTTTAA TGCAGCACTATTAGAGACTGTCGTACTACTGGAGACTGCAGCATTAATTTTCGAACGGCTACTGTCAATTATAGATCACTAGTATTTAGTCACAAAAGCTAATTTTTTAAGCAGAAATTCATAAAAATGTTTTCAATATTGCGAACTTTT GTAACAAAAAGACCCAGTAATTCAATTACTTTCGTAAATTATCAAAAAATCATCAAAAATATACAAAAAAATACCAAAAAATATTGAAACTTTCAAGTGACTCTTTCAATAGAAAATGGGGTGCAGCACTAATAGAGACTGCTGCACTAT TTTTCGGACCCTTTTTGAATGCAGCACTATTAGAGACTGCAGTATTTACTACTGGAGATGCAGCACTAATAGAGAATATACGGTATATACGTAATATATTCTTGCAGAAAAAAGTACGATTATCAATGAAAAATAGCTGATAAGAGGCTT TTGTTTGAACTAACAGACGGAACGACTCCGGTTTAGTTCAAAAAATTCTAAAAACACGTTGTGTCAGGCTGTCTCATTGCGGTTTGATCTACGAAAAATGCGGGAATATTTTTCCAGAAAAATTGTGACGTCAGCACGCTCTTAACCATG CGAAACGAGATGAGATGTCTGCGTCTCTTTTCCCGCATTTTTCGAAGATCAAAACGAATGGGACTTTCTGACTCCACGTGTAAAAAGGGGTTACGACGGACCCTGGCCTAGAAATTAGGCGTGAAAATTCTCGGGCACTGGATGTAGTGA ACGCCCGCGATGAAAAATTGGGGGAAAATTAGGCTTTCTTTGCGAGAAAGATTAATTAAAAATGTTTTCCTTTGTCGAAAATAATTTTTAAAAAACACACCACGTGTATTCAGCTCGACCAACGCCTCGAAAATTTTCAAAAAAGGCGGG AAAAATTAGTTGAATTCGCCAAGAGGAATTTCACCGCAGCGCGTGCAAAAATTTCAGCATTTGCGCGTGACGGTGTTTGCACAAATTACACCGAATGGTCGAGCTGAAAACACGTGCACACTTTTAAATAAAACTAGAAAATAAATCCCA GGCCTGCAAATATTGCACACAAAACCGTAATCCCCTTCGCGCTAAACAACACGCGCAACGATGCTCCGCTTGGGGACAAGGAAAAATTAATTTAACTCGGGATTTTCATTAAAAAATTAGGTTTTTAGTTAATTTTTCGATGTTTTCACT GCGAAAAAGTGTTAAAATAACGATTTTTCAACCTATTTTCAATTAATCCGTGCAAAAAATCGTGTATTTCTCGAGTTTTGAAAGAAATTTATGAAAATCGGCATTTTTAATAATGGTTTTTCAAATAAAAATATAATTTTTCGGTGCAGA AAAGTCGTTGCTCGTACAGTTTTTTTAAAGCATTTTCACATCAAAATCCTCCATTTTTCCAGTAAATCGATATGGAGTGCGACGAGACAAAGCTGAGCGACGGCGCAAGCGGCTGGGTGCCGAGTATCCCGACAGATATCGATTCAAAAG ACACACCGTTGCTCGATATATCTTCTCAGGCGATTTGGGCGCTTTCCAGTTGTAAAAGCGGTAAATTTTCCGACTTTCAAGGGAGAAAAGTGTAGAAAAATCGAAATTACTTCTTAAAAATCTCGTAAAAATCGAATTCTTTCAGGATTC GGCATCGACGAGCTCCTATCCGACAGTGTTGAGAAATATTGGCAAAGCGATGGCCCGCAGCCGCACACGATTCTTCTAGAATTCCAGAAAAAGACCGACGTGGCTATGATGATGTTCTATTTGGATTTTAAAAACGACGAGTCTTATACA CCGTCAAAGTTAGCATTTTTGGCTTTTTCAAACGAAAAAATACAATGAAACACTGAATATCTAGTTTTTTTCTCAATTTTTGCCTAAAAAACGGCGATTTTTCACTAGCTTTTCAATTAAAATTTGAACAAAAAGTTTTTTAAAGGAAAA ACATGAATTTCTAGCTTTTTCAGAGGTTTTCTATTAAAAAATAGAGATTTTTGTGATATCTGACTGAAAAATTACCAAACTGTCGATTTTTTTAAACTATTTTTCACTTAAAATCTGCAATTTTTTTTTTCGAGGAAACATGTGAATTTC AAGCTTTTTCAGAGATTTTCTATGAAAAAGGTTCGTGCCGAGACCCATGTGCTTTTAAACTTCAGAATTTTCCCAATTTTGAAATTAAAAAGAGAATGAAAATTGATTTTCATGGAAAAATGCGTTTTTGGCCCAAAACCTCCAAAAAGT ACAAATATAGGTCGACTTTCAACTGTTTTAGATCAATTTTTTTGCAGAATTCAAGTAAAAATGGGTTCATCTCACCAGGATATATTTTTCCGTCAAACACAAACATTCAACGAGCCCCAGGGATGGACATTTATCGATTTACGCGACAAA AATGGGAAACCGAATCGCGTTTTTTGGCTTCAAGTACAAGTTATTCAGAATCATCAAAATGGGAGAGATACTCATATAAGGTAGAGGAATTGAGAATTTCAGAACGAAAATTGCCGAAAAAATGAAATTTTAGCGAATTTGAGTCGGAAA TTTCGAAATTTGATTGATTTTAAGCAAATTTCCAACTAAAATCTTGAAAATTTGATCTTTTTAGATAAATTTTTTTTTAATTTTGTGCTTTTCAAAAAACCTCAAAAAACAATTAAAAATTGAAGTAAAATTAATTTTTCAACAATTTTT GAAAGGCCGAATTTTTGATTGAAAATTTTCACAATTTGTCCATTTTGTGGTGGGGCTTATTCCGAAAAATCGTTGTTTTTTTTTTCAAAAAAGTTATAAAAACTTTAAAATTGCCATGTAAAATATGTTTATTCTCAGACCTCGTAGGCA CGAAGCAGGCGTAGGTCGCCTCGCAATAAATTTGAAAATCTCAAGAAAAATCAATAAATTTGTGATTAATCAAAAAAATTTAATTTCCTGGTCCCAGCACGAATGCTATTTTTCGAAAAAAAAAAAGAGGCGAGCCTAATATAGACCACG CCCACAAAATGGGCAAAAGTTTGATTTTTCAAAAAATCGAAACAAAAATTTTTCCAATTTTGTGAGATTTTAAAATTTCCGGTTTTTGGAAAATCGAAAAAAAATTTCTCGTTTTTTAATTTTCAAAAAAAATTGTGCCTAAAATTCAAA AAAAAAATCAATACTTTCTCAAAATTTCCAGAAAACAGTCCATTTTCCAGGCACGTTCGAGTCCTTGGACCCCAGCGATCTCGTGTCTCCACAACGAATCGAATATTCACCGGAGAACCACACGGACCGATTCCCGATAAAAATATCACT AATTTCGACGACGAGGATTTTGCCAATTTTATCGATCACTCACTTGTTCACTTATCACTTCGTTAAATTTACCTCCAGTGATTCCAGATAATGAGCCAGTTTTGCATTGAAATTTAGTGCCAAAATATAGAAAATCGCATGATTTAACAT AAAATAGCGTTTCGAATTGAAACAATGGAAAAAAAGTGCTATGATGATTTTTTAACACTTTTAATTGTTCCAATTTGAAGTAAAATCTATTTTCAGATAAATCAACTGATTTTCTATATTCTGCCACTAAAGCTTAAAAACTTGCCCTGC TGTCCTAACCTTCAAATTGTTCCCTGCAAATTTTATTATTCTTGTTTCATATTTTTGCGATTGCTTCGCGAGACCCAAACTCACACATTTACCTGTAAAATATAATCGAATAATTATTTATATATTTTCTGTAAATTTCCTTAGTATACT ATAAATTTTCTGATCTCTCTTCAAAAATCGCTAGAAAAAATAAACAAATGTCGGTTTAAAAATTCCTGGTAATTTACCTTCTATAGAAAATTTTTCGAAAAAAAAACCGAAGAAATTCAGATGGAAATTCCCGATCCCGAACTGCCGGGA ATACCGATTGATCCGCAAGATTTGGAGATTCTAGACACGCCCACACGGTTTTACGAGAAGCTTTTAGTGCGTTTTTCGTGTCGGGACCCGGAAATTTGACATTTTTGGCGCGCGGCTTGTTAGACTCCAAACCTTTTCAAAGATTTTTTT TTCGAATTAAATAACATTCGTGCTTGGGCCCGGAAATTGAATTTTTGATTTGAAAACAATTTTTTTTGAGTCCAAAATTTTCAAAGTTTGTCCATTTTTGGCGCGTGGCCTAGTAGGATCCGCCCCTTCTAAATTTTTTTTGAGCAAGTT TTCTGAAGCATTGATTTCAAAAATTTTTTTTGGAAATTTCTGGTTTATTTTTCCGGTTTTTTTCCGAGTTGCTGTTTAAGTTTGGAGAAATTCCAGAATTTGTCAATTTTTGGGGCGTGGCTTTTTCAGTAAGCACAGTTTTTTTTTTTT GAAAAATTGAAATTTTCGCGGTGCGGTTCAAGAAAAACCACAAAAACTCAATGATTTTTTAACGAAAATTTCAAATTTCTTGCAAGACCTACTGCAATTTCGATTTTTAGAAACTTTTTGAAAAAAATCCGAATTTTCTGATTTAGCCCC GCCCCAAAAATGGAAAGATTTCCGAAAATTCGAACCAAAAGTTCGCAAAAACTTGAATTTCTCTCACACAGATTGACGCGCTAATTTGAATTTTTCCAAAAATAAGCCCCGCCCCAAAAATGGACAAATTTTAAAAATTTTGAACCAAAT AAATTCAATTTTTTTTCGCTTTTTTCCGTTTTCGAACAAAAAATTCTAAAAATATATGGTTCTAGGCGGGGCTCAGGCACCCATCTACCTACTTAAAAATGCGTTAAATTTCAGGAATTAACTGCATCAACCGAACGGCGTCTCGCATTG TGTAGTCTGTATTTGGGCGAAGGAGATCTCGAAAAAAATCTGATCGCTGCGATCCGAGAAAGATCCGAAAAATCCGAGATTGAAGTGACGATTCTGTTGGATTTTTTGCGCGGAACACGGACCAATTCAAGCGGCGAAAGTAGTGTAACA GTGCTGAAACCTATTTCGGAAAAGTCAAAAGTTGGTTTTTTTTGCAAAAAAAAATCGATAAATCGATAAAAACCGACAATTTTGAGAATTTTCATTTCAAATTTGAGTCCCACATGCGCCTTTAAATATGGTGTACTGTAGTTTTAGCTC GAATGTTGAATTTCAAAAATTGAGAATAAAGAAATGTCGTGACGAGACCCACAAATGTTTTGAAAAAAATTTTCAATTTCAAAAAAATGTAAAAAATTGGGAATTTCCCTCCAAAAGTTAAATTGGTTTAGTCACAAACTTTGAAATTTT GAAATAAAATTTTTTTCGGCTAAAAATAAGTATTTTTTAAAAACTATTTTGAAGAAAAAAAGTTAGGTCTCGCCACGATGTATCTTGTATATGTGTATCTAAATTGCCATGTCGTGACGAGACCCTCTCATATTTTACACTGCAACTTTT TCCTCACGAGGGACGAGGAAAAGTGGTTTCTAGGCCATGGCCGAGGGGCCGACAAGTTTCATCGGCCATTTATCTTGCTTTGTTTTCCGCCTGTTTTCTTTCGTTTTTCACAGCTTTTTCCCATTTTTTCTTATTAAAACTGATAAATAA ATATTTTTGCAGATGCCAAAACGATTTTCAAGTAAAAAAATCATGTATTCAGTGGGCAAGCAGCGGTGAAAGTGGGCATTGTAATATGATGGATTACGGGAATACAAAACCTAAACTTTTTCTGAAACATGATACATATGATGCTTAAAT GCTGAGACTACCTGATTTTCATAACGAGACCGCTGAAAAAGTTTTGAGGTTTTCAAAATTCAACTTTTTGTGCGAAAATCTCGACTTTTTCACCGAAAAAGTTGAATTTTGGAAACCTCAAAACTTTTTCAGCGGTCTTGATATGAAAAT CAGGTAGCTTCAGCATCTAAGCAGCATATGTATCATGTTAAAGAAAAAGTTTAGGTTTTGTATTCCTGTAATCCATCATATTACATTGCCCACTTTCACCGCTGCTTGCCCACTGAATACATAATTTTTTCACTTGGAAATTGTTTTAGC Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
11 Example: C. elegans (I: 43,500-52,050) GAAGAAATGGAGCATTTGCGCTCCATCACACTCTCAGACAATTTCATTTTCCACATCCTATATATATTTTGGTTTTTCTGTCGTATTTTGTTTTAATTTATTGGTATTTCGTTCAAAAATAATTATTTTGACTGTATTTTTGGTTGCATA CATGTAGAACTGCTGTTTTTTAAGATATTCTGCCCATTCAAGTTTTTCAGTGTAAAATTGATATATTTCATTCCAACTGAAAATGAGATCGAAACGATGGAAAACCTCGGATATTACTGATTATGGAAAGAAGAGAAAAGAATCGGAAAG TTGTGGATCAAGTTCACCGATTCTCGAAACACAGTCATCTGGCGGTGCGGAACTTGACGAAGTTACTGAGGATGAATATTCTAGTAATTCGAGCAGTAATGAAACTAGCGACGAAGAGGAAAACTCAGAAGTACCAAATGTCTTATCTAT AACAGAAAGAGGTAAGAATTGCGTCTTCTAGTGATCATACTTTTCGCCAGATTCCCTAATGTAATATATTTTGTTGTAGAGAAAAGTTGGCAAAAGTTAACGGAAAACGATTTGGGACGAATTCGTTTCATCTTGAAGTACACTAGCAAT ACTAAAAAATGCGTGAACGAGTATTTTCAATATAATCATGGGCAAAACAATGAAATTATGAAAAGTCTATTATTGGATACCGATGGAACTATGACTGCAAAGGCTTGTTCGGAATGTGCCTACGATTTGAATCAGTAAGTTACTCTCTCG ATTTATTCCCAAAATTAATATGTGCTTCAGGTGCCACTGCAAAAAACCGCTTCGCTTCATCAATGCTCCGTGTGGTTGGTTTGCTATTCAAAACTATAAATAGTTCACTGTTTCCGTTCAGAGGTCATCAACCAAGTTCTTCATGTTGAA AATGCGGAGCCCACCAGGATCAACCATGTAATCGCAACACTCTTCCGGAATCACATTGGCGAGATTTTGTTGGTCCACTCTATTTCTGTGCGAGAACTGTGATAAAACTAGTATTTTCAGCACAAAGGCTCGAACTGCGGAAGCTCGCGC ATCTGAAGAAGCTCAAATCAGGATTCAAATCCAAGACAACTCGAACGCATTCCAAAGATCGTATCATAACGATCCACAACCTTCATCAGCCGAAGAACATGAGGAAGATATCGTGGTGGATGGCTGAGTACGGAGCTCAAATGCCTTAAG GCGAAACAATTGGTTTTTTAATTTGCTGGTTATCATGTTAGATTTTGAACGTGTTAGGTCTTTCAATTGTTTTTTTTTTTCGAAATGTTGTTGTTCTAATAAATTTGTTTTATTTAATCAAACGTTTTTTAGTCTACTACGGGCGTGAAG CCAGATATCAGTGGTATCTTCTTATCAGAAGCTGAATCATTTCCGGTTGACAATGTTTGAAGGACATAAGAAAGGCTGTGTTACTGATTTCGACCATTGATTTGTTTATATATGGATATGTTCCACTGCCTTTTGGAAAGGCAGTATTCC CGGTATATATGGGCCTAATACGGAATCTAAAATAACCTGACACAAACCTGACGTTGACCTGTTGCCGGCCCGCGGCGGCTTAGTGTCAACTTGACAGCGGGTCGCGATTTCACCTGCCAGTTGTTCTCCATTCAGCAGCCAGCGACCTGC TGGCAGGTTGCCACTAACCTGACGCGGTTTACCTGTGTTATCGGCGCGTGCATAGCTTAGTGGTTTCAGGAAATGATGCTAGTAATCAGAAGATCGGGGTTCGGGAAACGGCAGGGGCTTGAAGGTTAGGTTCTATGAAGCAGGGCGAAG GGTTGACAAGGAGAGGCAATAAGCAAGTAGTAGGGGTTCTCTAGAAAACATTTTTGTCTTTAATATGCGTTTCCTACTGATTTATTATTGATATTTGGATCCCCTTTTCTAGAAAAAAAAATCAGAATCAGCAGAAAAATTTGAGAAAAA GTCATAGCAAATCAGAGTTGGTCAGAGTAAATCAGAGCTAGTCATAGTAAATCATAGCTAGTCAGAGAATATCAGAGTTAATCAGGGTAATAAGTAGACCTAGTCATAGTAAATCAGAGCTAGGCATAGTAAAGCGTGGTTACTCCGAGT AAAACCACACTTGCACCGAACTGCGGTTAGTGTGCTTTACCATTATGTAACTCCGCTTTTTACTCTGAGTTAGTATGATATGGTTTGTCTGAGCTGTGGTTGGGCTTCGCGGGAAACTTGAATAATTCGAGACAAAATCTAATTTTAGCG AATTTTCTTTAATTTCTTTGAGGTTTCTACGACAGAACTCGAAAAATTTCGGGTTTTAATGTTTACACATTTTATTTAAAATTGAATAATCAACTGCGGGACTCCTCGAAAATCACATGCTCATTTAAATTTTGAAGTTCAAACCTCAAA AAACGCGCAAAAACCAAATTCAGCTAGGATATCAAATTTATGATTGAAATCTATATTTTGATGCGGTGTTTCTGAAGTTTTCGCGATAAAATCCGAATAATAATTCCACGTACCGTATATTCTCTATCTAATTTCCAGGTCATTTTTTAA TGCAGCACTATTAGAGACTGTCGTACTACTGGAGACTGCAGCATTAATTTTCGAACGGCTACTGTCAATTATAGATCACTAGTATTTAGTCACAAAAGCTAATTTTTTAAGCAGAAATTCATAAAAATGTTTTCAATATTGCGAACTTTT GTAACAAAAAGACCCAGTAATTCAATTACTTTCGTAAATTATCAAAAAATCATCAAAAATATACAAAAAAATACCAAAAAATATTGAAACTTTCAAGTGACTCTTTCAATAGAAAATGGGGTGCAGCACTAATAGAGACTGCTGCACTAT TTTTCGGACCCTTTTTGAATGCAGCACTATTAGAGACTGCAGTATTTACTACTGGAGATGCAGCACTAATAGAGAATATACGGTATATACGTAATATATTCTTGCAGAAAAAAGTACGATTATCAATGAAAAATAGCTGATAAGAGGCTT TTGTTTGAACTAACAGACGGAACGACTCCGGTTTAGTTCAAAAAATTCTAAAAACACGTTGTGTCAGGCTGTCTCATTGCGGTTTGATCTACGAAAAATGCGGGAATATTTTTCCAGAAAAATTGTGACGTCAGCACGCTCTTAACCATG CGAAACGAGATGAGATGTCTGCGTCTCTTTTCCCGCATTTTTCGAAGATCAAAACGAATGGGACTTTCTGACTCCACGTGTAAAAAGGGGTTACGACGGACCCTGGCCTAGAAATTAGGCGTGAAAATTCTCGGGCACTGGATGTAGTGA ACGCCCGCGATGAAAAATTGGGGGAAAATTAGGCTTTCTTTGCGAGAAAGATTAATTAAAAATGTTTTCCTTTGTCGAAAATAATTTTTAAAAAACACACCACGTGTATTCAGCTCGACCAACGCCTCGAAAATTTTCAAAAAAGGCGGG AAAAATTAGTTGAATTCGCCAAGAGGAATTTCACCGCAGCGCGTGCAAAAATTTCAGCATTTGCGCGTGACGGTGTTTGCACAAATTACACCGAATGGTCGAGCTGAAAACACGTGCACACTTTTAAATAAAACTAGAAAATAAATCCCA GGCCTGCAAATATTGCACACAAAACCGTAATCCCCTTCGCGCTAAACAACACGCGCAACGATGCTCCGCTTGGGGACAAGGAAAAATTAATTTAACTCGGGATTTTCATTAAAAAATTAGGTTTTTAGTTAATTTTTCGATGTTTTCACT GCGAAAAAGTGTTAAAATAACGATTTTTCAACCTATTTTCAATTAATCCGTGCAAAAAATCGTGTATTTCTCGAGTTTTGAAAGAAATTTATGAAAATCGGCATTTTTAATAATGGTTTTTCAAATAAAAATATAATTTTTCGGTGCAGA AAAGTCGTTGCTCGTACAGTTTTTTTAAAGCATTTTCACATCAAAATCCTCCATTTTTCCAGTAAATCGATATGGAGTGCGACGAGACAAAGCTGAGCGACGGCGCAAGCGGCTGGGTGCCGAGTATCCCGACAGATATCGATTCAAAAG ACACACCGTTGCTCGATATATCTTCTCAGGCGATTTGGGCGCTTTCCAGTTGTAAAAGCGGTAAATTTTCCGACTTTCAAGGGAGAAAAGTGTAGAAAAATCGAAATTACTTCTTAAAAATCTCGTAAAAATCGAATTCTTTCAGGATTC GGCATCGACGAGCTCCTATCCGACAGTGTTGAGAAATATTGGCAAAGCGATGGCCCGCAGCCGCACACGATTCTTCTAGAATTCCAGAAAAAGACCGACGTGGCTATGATGATGTTCTATTTGGATTTTAAAAACGACGAGTCTTATACA CCGTCAAAGTTAGCATTTTTGGCTTTTTCAAACGAAAAAATACAATGAAACACTGAATATCTAGTTTTTTTCTCAATTTTTGCCTAAAAAACGGCGATTTTTCACTAGCTTTTCAATTAAAATTTGAACAAAAAGTTTTTTAAAGGAAAA ACATGAATTTCTAGCTTTTTCAGAGGTTTTCTATTAAAAAATAGAGATTTTTGTGATATCTGACTGAAAAATTACCAAACTGTCGATTTTTTTAAACTATTTTTCACTTAAAATCTGCAATTTTTTTTTTCGAGGAAACATGTGAATTTC AAGCTTTTTCAGAGATTTTCTATGAAAAAGGTTCGTGCCGAGACCCATGTGCTTTTAAACTTCAGAATTTTCCCAATTTTGAAATTAAAAAGAGAATGAAAATTGATTTTCATGGAAAAATGCGTTTTTGGCCCAAAACCTCCAAAAAGT ACAAATATAGGTCGACTTTCAACTGTTTTAGATCAATTTTTTTGCAGAATTCAAGTAAAAATGGGTTCATCTCACCAGGATATATTTTTCCGTCAAACACAAACATTCAACGAGCCCCAGGGATGGACATTTATCGATTTACGCGACAAA AATGGGAAACCGAATCGCGTTTTTTGGCTTCAAGTACAAGTTATTCAGAATCATCAAAATGGGAGAGATACTCATATAAGGTAGAGGAATTGAGAATTTCAGAACGAAAATTGCCGAAAAAATGAAATTTTAGCGAATTTGAGTCGGAAA TTTCGAAATTTGATTGATTTTAAGCAAATTTCCAACTAAAATCTTGAAAATTTGATCTTTTTAGATAAATTTTTTTTTAATTTTGTGCTTTTCAAAAAACCTCAAAAAACAATTAAAAATTGAAGTAAAATTAATTTTTCAACAATTTTT GAAAGGCCGAATTTTTGATTGAAAATTTTCACAATTTGTCCATTTTGTGGTGGGGCTTATTCCGAAAAATCGTTGTTTTTTTTTTCAAAAAAGTTATAAAAACTTTAAAATTGCCATGTAAAATATGTTTATTCTCAGACCTCGTAGGCA CGAAGCAGGCGTAGGTCGCCTCGCAATAAATTTGAAAATCTCAAGAAAAATCAATAAATTTGTGATTAATCAAAAAAATTTAATTTCCTGGTCCCAGCACGAATGCTATTTTTCGAAAAAAAAAAAGAGGCGAGCCTAATATAGACCACG CCCACAAAATGGGCAAAAGTTTGATTTTTCAAAAAATCGAAACAAAAATTTTTCCAATTTTGTGAGATTTTAAAATTTCCGGTTTTTGGAAAATCGAAAAAAAATTTCTCGTTTTTTAATTTTCAAAAAAAATTGTGCCTAAAATTCAAA AAAAAAATCAATACTTTCTCAAAATTTCCAGAAAACAGTCCATTTTCCAGGCACGTTCGAGTCCTTGGACCCCAGCGATCTCGTGTCTCCACAACGAATCGAATATTCACCGGAGAACCACACGGACCGATTCCCGATAAAAATATCACT AATTTCGACGACGAGGATTTTGCCAATTTTATCGATCACTCACTTGTTCACTTATCACTTCGTTAAATTTACCTCCAGTGATTCCAGATAATGAGCCAGTTTTGCATTGAAATTTAGTGCCAAAATATAGAAAATCGCATGATTTAACAT AAAATAGCGTTTCGAATTGAAACAATGGAAAAAAAGTGCTATGATGATTTTTTAACACTTTTAATTGTTCCAATTTGAAGTAAAATCTATTTTCAGATAAATCAACTGATTTTCTATATTCTGCCACTAAAGCTTAAAAACTTGCCCTGC TGTCCTAACCTTCAAATTGTTCCCTGCAAATTTTATTATTCTTGTTTCATATTTTTGCGATTGCTTCGCGAGACCCAAACTCACACATTTACCTGTAAAATATAATCGAATAATTATTTATATATTTTCTGTAAATTTCCTTAGTATACT ATAAATTTTCTGATCTCTCTTCAAAAATCGCTAGAAAAAATAAACAAATGTCGGTTTAAAAATTCCTGGTAATTTACCTTCTATAGAAAATTTTTCGAAAAAAAAACCGAAGAAATTCAGATGGAAATTCCCGATCCCGAACTGCCGGGA ATACCGATTGATCCGCAAGATTTGGAGATTCTAGACACGCCCACACGGTTTTACGAGAAGCTTTTAGTGCGTTTTTCGTGTCGGGACCCGGAAATTTGACATTTTTGGCGCGCGGCTTGTTAGACTCCAAACCTTTTCAAAGATTTTTTT TTCGAATTAAATAACATTCGTGCTTGGGCCCGGAAATTGAATTTTTGATTTGAAAACAATTTTTTTTGAGTCCAAAATTTTCAAAGTTTGTCCATTTTTGGCGCGTGGCCTAGTAGGATCCGCCCCTTCTAAATTTTTTTTGAGCAAGTT TTCTGAAGCATTGATTTCAAAAATTTTTTTTGGAAATTTCTGGTTTATTTTTCCGGTTTTTTTCCGAGTTGCTGTTTAAGTTTGGAGAAATTCCAGAATTTGTCAATTTTTGGGGCGTGGCTTTTTCAGTAAGCACAGTTTTTTTTTTTT GAAAAATTGAAATTTTCGCGGTGCGGTTCAAGAAAAACCACAAAAACTCAATGATTTTTTAACGAAAATTTCAAATTTCTTGCAAGACCTACTGCAATTTCGATTTTTAGAAACTTTTTGAAAAAAATCCGAATTTTCTGATTTAGCCCC GCCCCAAAAATGGAAAGATTTCCGAAAATTCGAACCAAAAGTTCGCAAAAACTTGAATTTCTCTCACACAGATTGACGCGCTAATTTGAATTTTTCCAAAAATAAGCCCCGCCCCAAAAATGGACAAATTTTAAAAATTTTGAACCAAAT AAATTCAATTTTTTTTCGCTTTTTTCCGTTTTCGAACAAAAAATTCTAAAAATATATGGTTCTAGGCGGGGCTCAGGCACCCATCTACCTACTTAAAAATGCGTTAAATTTCAGGAATTAACTGCATCAACCGAACGGCGTCTCGCATTG TGTAGTCTGTATTTGGGCGAAGGAGATCTCGAAAAAAATCTGATCGCTGCGATCCGAGAAAGATCCGAAAAATCCGAGATTGAAGTGACGATTCTGTTGGATTTTTTGCGCGGAACACGGACCAATTCAAGCGGCGAAAGTAGTGTAACA GTGCTGAAACCTATTTCGGAAAAGTCAAAAGTTGGTTTTTTTTGCAAAAAAAAATCGATAAATCGATAAAAACCGACAATTTTGAGAATTTTCATTTCAAATTTGAGTCCCACATGCGCCTTTAAATATGGTGTACTGTAGTTTTAGCTC GAATGTTGAATTTCAAAAATTGAGAATAAAGAAATGTCGTGACGAGACCCACAAATGTTTTGAAAAAAATTTTCAATTTCAAAAAAATGTAAAAAATTGGGAATTTCCCTCCAAAAGTTAAATTGGTTTAGTCACAAACTTTGAAATTTT GAAATAAAATTTTTTTCGGCTAAAAATAAGTATTTTTTAAAAACTATTTTGAAGAAAAAAAGTTAGGTCTCGCCACGATGTATCTTGTATATGTGTATCTAAATTGCCATGTCGTGACGAGACCCTCTCATATTTTACACTGCAACTTTT TCCTCACGAGGGACGAGGAAAAGTGGTTTCTAGGCCATGGCCGAGGGGCCGACAAGTTTCATCGGCCATTTATCTTGCTTTGTTTTCCGCCTGTTTTCTTTCGTTTTTCACAGCTTTTTCCCATTTTTTCTTATTAAAACTGATAAATAA ATATTTTTGCAGATGCCAAAACGATTTTCAAGTAAAAAAATCATGTATTCAGTGGGCAAGCAGCGGTGAAAGTGGGCATTGTAATATGATGGATTACGGGAATACAAAACCTAAACTTTTTCTGAAACATGATACATATGATGCTTAAAT GCTGAGACTACCTGATTTTCATAACGAGACCGCTGAAAAAGTTTTGAGGTTTTCAAAATTCAACTTTTTGTGCGAAAATCTCGACTTTTTCACCGAAAAAGTTGAATTTTGGAAACCTCAAAACTTTTTCAGCGGTCTTGATATGAAAAT CAGGTAGCTTCAGCATCTAAGCAGCATATGTATCATGTTAAAGAAAAAGTTTAGGTTTTGTATTCCTGTAATCCATCATATTACATTGCCCACTTTCACCGCTGCTTGCCCACTGAATACATAATTTTTTCACTTGGAAATTGTTTTAGC Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
12 Example: C. elegans (I: 43,500-52,050) GAAGAAATGGAGCATTTGCGCTCCATCACACTCTCAGACAATTTCATTTTCCACATCCTATATATATTTTGGTTTTTCTGTCGTATTTTGTTTTAATTTATTGGTATTTCGTTCAAAAATAATTATTTTGACTGTATTTTTGGTTGCATA CATGTAGAACTGCTGTTTTTTAAGATATTCTGCCCATTCAAGTTTTTCAGTGTAAAATTGATATATTTCATTCCAACTGAAAATGAGATCGAAACGATGGAAAACCTCGGATATTACTGATTATGGAAAGAAGAGAAAAGAATCGGAAAG TTGTGGATCAAGTTCACCGATTCTCGAAACACAGTCATCTGGCGGTGCGGAACTTGACGAAGTTACTGAGGATGAATATTCTAGTAATTCGAGCAGTAATGAAACTAGCGACGAAGAGGAAAACTCAGAAGTACCAAATGTCTTATCTAT AACAGAAAGAGGTAAGAATTGCGTCTTCTAGTGATCATACTTTTCGCCAGATTCCCTAATGTAATATATTTTGTTGTAGAGAAAAGTTGGCAAAAGTTAACGGAAAACGATTTGGGACGAATTCGTTTCATCTTGAAGTACACTAGCAAT ACTAAAAAATGCGTGAACGAGTATTTTCAATATAATCATGGGCAAAACAATGAAATTATGAAAAGTCTATTATTGGATACCGATGGAACTATGACTGCAAAGGCTTGTTCGGAATGTGCCTACGATTTGAATCAGTAAGTTACTCTCTCG ATTTATTCCCAAAATTAATATGTGCTTCAGGTGCCACTGCAAAAAACCGCTTCGCTTCATCAATGCTCCGTGTGGTTGGTTTGCTATTCAAAACTATAAATAGTTCACTGTTTCCGTTCAGAGGTCATCAACCAAGTTCTTCATGTTGAA AATGCGGAGCCCACCAGGATCAACCATGTAATCGCAACACTCTTCCGGAATCACATTGGCGAGATTTTGTTGGTCCACTCTATTTCTGTGCGAGAACTGTGATAAAACTAGTATTTTCAGCACAAAGGCTCGAACTGCGGAAGCTCGCGC ATCTGAAGAAGCTCAAATCAGGATTCAAATCCAAGACAACTCGAACGCATTCCAAAGATCGTATCATAACGATCCACAACCTTCATCAGCCGAAGAACATGAGGAAGATATCGTGGTGGATGGCTGAGTACGGAGCTCAAATGCCTTAAG GCGAAACAATTGGTTTTTTAATTTGCTGGTTATCATGTTAGATTTTGAACGTGTTAGGTCTTTCAATTGTTTTTTTTTTTCGAAATGTTGTTGTTCTAATAAATTTGTTTTATTTAATCAAACGTTTTTTAGTCTACTACGGGCGTGAAG CCAGATATCAGTGGTATCTTCTTATCAGAAGCTGAATCATTTCCGGTTGACAATGTTTGAAGGACATAAGAAAGGCTGTGTTACTGATTTCGACCATTGATTTGTTTATATATGGATATGTTCCACTGCCTTTTGGAAAGGCAGTATTCC CGGTATATATGGGCCTAATACGGAATCTAAAATAACCTGACACAAACCTGACGTTGACCTGTTGCCGGCCCGCGGCGGCTTAGTGTCAACTTGACAGCGGGTCGCGATTTCACCTGCCAGTTGTTCTCCATTCAGCAGCCAGCGACCTGC TGGCAGGTTGCCACTAACCTGACGCGGTTTACCTGTGTTATCGGCGCGTGCATAGCTTAGTGGTTTCAGGAAATGATGCTAGTAATCAGAAGATCGGGGTTCGGGAAACGGCAGGGGCTTGAAGGTTAGGTTCTATGAAGCAGGGCGAAG GGTTGACAAGGAGAGGCAATAAGCAAGTAGTAGGGGTTCTCTAGAAAACATTTTTGTCTTTAATATGCGTTTCCTACTGATTTATTATTGATATTTGGATCCCCTTTTCTAGAAAAAAAAATCAGAATCAGCAGAAAAATTTGAGAAAAA GTCATAGCAAATCAGAGTTGGTCAGAGTAAATCAGAGCTAGTCATAGTAAATCATAGCTAGTCAGAGAATATCAGAGTTAATCAGGGTAATAAGTAGACCTAGTCATAGTAAATCAGAGCTAGGCATAGTAAAGCGTGGTTACTCCGAGT AAAACCACACTTGCACCGAACTGCGGTTAGTGTGCTTTACCATTATGTAACTCCGCTTTTTACTCTGAGTTAGTATGATATGGTTTGTCTGAGCTGTGGTTGGGCTTCGCGGGAAACTTGAATAATTCGAGACAAAATCTAATTTTAGCG AATTTTCTTTAATTTCTTTGAGGTTTCTACGACAGAACTCGAAAAATTTCGGGTTTTAATGTTTACACATTTTATTTAAAATTGAATAATCAACTGCGGGACTCCTCGAAAATCACATGCTCATTTAAATTTTGAAGTTCAAACCTCAAA AAACGCGCAAAAACCAAATTCAGCTAGGATATCAAATTTATGATTGAAATCTATATTTTGATGCGGTGTTTCTGAAGTTTTCGCGATAAAATCCGAATAATAATTCCACGTACCGTATATTCTCTATCTAATTTCCAGGTCATTTTTTAA TGCAGCACTATTAGAGACTGTCGTACTACTGGAGACTGCAGCATTAATTTTCGAACGGCTACTGTCAATTATAGATCACTAGTATTTAGTCACAAAAGCTAATTTTTTAAGCAGAAATTCATAAAAATGTTTTCAATATTGCGAACTTTT GTAACAAAAAGACCCAGTAATTCAATTACTTTCGTAAATTATCAAAAAATCATCAAAAATATACAAAAAAATACCAAAAAATATTGAAACTTTCAAGTGACTCTTTCAATAGAAAATGGGGTGCAGCACTAATAGAGACTGCTGCACTAT TTTTCGGACCCTTTTTGAATGCAGCACTATTAGAGACTGCAGTATTTACTACTGGAGATGCAGCACTAATAGAGAATATACGGTATATACGTAATATATTCTTGCAGAAAAAAGTACGATTATCAATGAAAAATAGCTGATAAGAGGCTT TTGTTTGAACTAACAGACGGAACGACTCCGGTTTAGTTCAAAAAATTCTAAAAACACGTTGTGTCAGGCTGTCTCATTGCGGTTTGATCTACGAAAAATGCGGGAATATTTTTCCAGAAAAATTGTGACGTCAGCACGCTCTTAACCATG CGAAACGAGATGAGATGTCTGCGTCTCTTTTCCCGCATTTTTCGAAGATCAAAACGAATGGGACTTTCTGACTCCACGTGTAAAAAGGGGTTACGACGGACCCTGGCCTAGAAATTAGGCGTGAAAATTCTCGGGCACTGGATGTAGTGA ACGCCCGCGATGAAAAATTGGGGGAAAATTAGGCTTTCTTTGCGAGAAAGATTAATTAAAAATGTTTTCCTTTGTCGAAAATAATTTTTAAAAAACACACCACGTGTATTCAGCTCGACCAACGCCTCGAAAATTTTCAAAAAAGGCGGG AAAAATTAGTTGAATTCGCCAAGAGGAATTTCACCGCAGCGCGTGCAAAAATTTCAGCATTTGCGCGTGACGGTGTTTGCACAAATTACACCGAATGGTCGAGCTGAAAACACGTGCACACTTTTAAATAAAACTAGAAAATAAATCCCA GGCCTGCAAATATTGCACACAAAACCGTAATCCCCTTCGCGCTAAACAACACGCGCAACGATGCTCCGCTTGGGGACAAGGAAAAATTAATTTAACTCGGGATTTTCATTAAAAAATTAGGTTTTTAGTTAATTTTTCGATGTTTTCACT GCGAAAAAGTGTTAAAATAACGATTTTTCAACCTATTTTCAATTAATCCGTGCAAAAAATCGTGTATTTCTCGAGTTTTGAAAGAAATTTATGAAAATCGGCATTTTTAATAATGGTTTTTCAAATAAAAATATAATTTTTCGGTGCAGA AAAGTCGTTGCTCGTACAGTTTTTTTAAAGCATTTTCACATCAAAATCCTCCATTTTTCCAGTAAATCGATATGGAGTGCGACGAGACAAAGCTGAGCGACGGCGCAAGCGGCTGGGTGCCGAGTATCCCGACAGATATCGATTCAAAAG ACACACCGTTGCTCGATATATCTTCTCAGGCGATTTGGGCGCTTTCCAGTTGTAAAAGCGGTAAATTTTCCGACTTTCAAGGGAGAAAAGTGTAGAAAAATCGAAATTACTTCTTAAAAATCTCGTAAAAATCGAATTCTTTCAGGATTC GGCATCGACGAGCTCCTATCCGACAGTGTTGAGAAATATTGGCAAAGCGATGGCCCGCAGCCGCACACGATTCTTCTAGAATTCCAGAAAAAGACCGACGTGGCTATGATGATGTTCTATTTGGATTTTAAAAACGACGAGTCTTATACA CCGTCAAAGTTAGCATTTTTGGCTTTTTCAAACGAAAAAATACAATGAAACACTGAATATCTAGTTTTTTTCTCAATTTTTGCCTAAAAAACGGCGATTTTTCACTAGCTTTTCAATTAAAATTTGAACAAAAAGTTTTTTAAAGGAAAA ACATGAATTTCTAGCTTTTTCAGAGGTTTTCTATTAAAAAATAGAGATTTTTGTGATATCTGACTGAAAAATTACCAAACTGTCGATTTTTTTAAACTATTTTTCACTTAAAATCTGCAATTTTTTTTTTCGAGGAAACATGTGAATTTC AAGCTTTTTCAGAGATTTTCTATGAAAAAGGTTCGTGCCGAGACCCATGTGCTTTTAAACTTCAGAATTTTCCCAATTTTGAAATTAAAAAGAGAATGAAAATTGATTTTCATGGAAAAATGCGTTTTTGGCCCAAAACCTCCAAAAAGT ACAAATATAGGTCGACTTTCAACTGTTTTAGATCAATTTTTTTGCAGAATTCAAGTAAAAATGGGTTCATCTCACCAGGATATATTTTTCCGTCAAACACAAACATTCAACGAGCCCCAGGGATGGACATTTATCGATTTACGCGACAAA AATGGGAAACCGAATCGCGTTTTTTGGCTTCAAGTACAAGTTATTCAGAATCATCAAAATGGGAGAGATACTCATATAAGGTAGAGGAATTGAGAATTTCAGAACGAAAATTGCCGAAAAAATGAAATTTTAGCGAATTTGAGTCGGAAA TTTCGAAATTTGATTGATTTTAAGCAAATTTCCAACTAAAATCTTGAAAATTTGATCTTTTTAGATAAATTTTTTTTTAATTTTGTGCTTTTCAAAAAACCTCAAAAAACAATTAAAAATTGAAGTAAAATTAATTTTTCAACAATTTTT GAAAGGCCGAATTTTTGATTGAAAATTTTCACAATTTGTCCATTTTGTGGTGGGGCTTATTCCGAAAAATCGTTGTTTTTTTTTTCAAAAAAGTTATAAAAACTTTAAAATTGCCATGTAAAATATGTTTATTCTCAGACCTCGTAGGCA CGAAGCAGGCGTAGGTCGCCTCGCAATAAATTTGAAAATCTCAAGAAAAATCAATAAATTTGTGATTAATCAAAAAAATTTAATTTCCTGGTCCCAGCACGAATGCTATTTTTCGAAAAAAAAAAAGAGGCGAGCCTAATATAGACCACG CCCACAAAATGGGCAAAAGTTTGATTTTTCAAAAAATCGAAACAAAAATTTTTCCAATTTTGTGAGATTTTAAAATTTCCGGTTTTTGGAAAATCGAAAAAAAATTTCTCGTTTTTTAATTTTCAAAAAAAATTGTGCCTAAAATTCAAA AAAAAAATCAATACTTTCTCAAAATTTCCAGAAAACAGTCCATTTTCCAGGCACGTTCGAGTCCTTGGACCCCAGCGATCTCGTGTCTCCACAACGAATCGAATATTCACCGGAGAACCACACGGACCGATTCCCGATAAAAATATCACT AATTTCGACGACGAGGATTTTGCCAATTTTATCGATCACTCACTTGTTCACTTATCACTTCGTTAAATTTACCTCCAGTGATTCCAGATAATGAGCCAGTTTTGCATTGAAATTTAGTGCCAAAATATAGAAAATCGCATGATTTAACAT AAAATAGCGTTTCGAATTGAAACAATGGAAAAAAAGTGCTATGATGATTTTTTAACACTTTTAATTGTTCCAATTTGAAGTAAAATCTATTTTCAGATAAATCAACTGATTTTCTATATTCTGCCACTAAAGCTTAAAAACTTGCCCTGC TGTCCTAACCTTCAAATTGTTCCCTGCAAATTTTATTATTCTTGTTTCATATTTTTGCGATTGCTTCGCGAGACCCAAACTCACACATTTACCTGTAAAATATAATCGAATAATTATTTATATATTTTCTGTAAATTTCCTTAGTATACT ATAAATTTTCTGATCTCTCTTCAAAAATCGCTAGAAAAAATAAACAAATGTCGGTTTAAAAATTCCTGGTAATTTACCTTCTATAGAAAATTTTTCGAAAAAAAAACCGAAGAAATTCAGATGGAAATTCCCGATCCCGAACTGCCGGGA ATACCGATTGATCCGCAAGATTTGGAGATTCTAGACACGCCCACACGGTTTTACGAGAAGCTTTTAGTGCGTTTTTCGTGTCGGGACCCGGAAATTTGACATTTTTGGCGCGCGGCTTGTTAGACTCCAAACCTTTTCAAAGATTTTTTT TTCGAATTAAATAACATTCGTGCTTGGGCCCGGAAATTGAATTTTTGATTTGAAAACAATTTTTTTTGAGTCCAAAATTTTCAAAGTTTGTCCATTTTTGGCGCGTGGCCTAGTAGGATCCGCCCCTTCTAAATTTTTTTTGAGCAAGTT TTCTGAAGCATTGATTTCAAAAATTTTTTTTGGAAATTTCTGGTTTATTTTTCCGGTTTTTTTCCGAGTTGCTGTTTAAGTTTGGAGAAATTCCAGAATTTGTCAATTTTTGGGGCGTGGCTTTTTCAGTAAGCACAGTTTTTTTTTTTT GAAAAATTGAAATTTTCGCGGTGCGGTTCAAGAAAAACCACAAAAACTCAATGATTTTTTAACGAAAATTTCAAATTTCTTGCAAGACCTACTGCAATTTCGATTTTTAGAAACTTTTTGAAAAAAATCCGAATTTTCTGATTTAGCCCC GCCCCAAAAATGGAAAGATTTCCGAAAATTCGAACCAAAAGTTCGCAAAAACTTGAATTTCTCTCACACAGATTGACGCGCTAATTTGAATTTTTCCAAAAATAAGCCCCGCCCCAAAAATGGACAAATTTTAAAAATTTTGAACCAAAT AAATTCAATTTTTTTTCGCTTTTTTCCGTTTTCGAACAAAAAATTCTAAAAATATATGGTTCTAGGCGGGGCTCAGGCACCCATCTACCTACTTAAAAATGCGTTAAATTTCAGGAATTAACTGCATCAACCGAACGGCGTCTCGCATTG TGTAGTCTGTATTTGGGCGAAGGAGATCTCGAAAAAAATCTGATCGCTGCGATCCGAGAAAGATCCGAAAAATCCGAGATTGAAGTGACGATTCTGTTGGATTTTTTGCGCGGAACACGGACCAATTCAAGCGGCGAAAGTAGTGTAACA GTGCTGAAACCTATTTCGGAAAAGTCAAAAGTTGGTTTTTTTTGCAAAAAAAAATCGATAAATCGATAAAAACCGACAATTTTGAGAATTTTCATTTCAAATTTGAGTCCCACATGCGCCTTTAAATATGGTGTACTGTAGTTTTAGCTC GAATGTTGAATTTCAAAAATTGAGAATAAAGAAATGTCGTGACGAGACCCACAAATGTTTTGAAAAAAATTTTCAATTTCAAAAAAATGTAAAAAATTGGGAATTTCCCTCCAAAAGTTAAATTGGTTTAGTCACAAACTTTGAAATTTT GAAATAAAATTTTTTTCGGCTAAAAATAAGTATTTTTTAAAAACTATTTTGAAGAAAAAAAGTTAGGTCTCGCCACGATGTATCTTGTATATGTGTATCTAAATTGCCATGTCGTGACGAGACCCTCTCATATTTTACACTGCAACTTTT TCCTCACGAGGGACGAGGAAAAGTGGTTTCTAGGCCATGGCCGAGGGGCCGACAAGTTTCATCGGCCATTTATCTTGCTTTGTTTTCCGCCTGTTTTCTTTCGTTTTTCACAGCTTTTTCCCATTTTTTCTTATTAAAACTGATAAATAA ATATTTTTGCAGATGCCAAAACGATTTTCAAGTAAAAAAATCATGTATTCAGTGGGCAAGCAGCGGTGAAAGTGGGCATTGTAATATGATGGATTACGGGAATACAAAACCTAAACTTTTTCTGAAACATGATACATATGATGCTTAAAT GCTGAGACTACCTGATTTTCATAACGAGACCGCTGAAAAAGTTTTGAGGTTTTCAAAATTCAACTTTTTGTGCGAAAATCTCGACTTTTTCACCGAAAAAGTTGAATTTTGGAAACCTCAAAACTTTTTCAGCGGTCTTGATATGAAAAT CAGGTAGCTTCAGCATCTAAGCAGCATATGTATCATGTTAAAGAAAAAGTTTAGGTTTTGTATTCCTGTAATCCATCATATTACATTGCCCACTTTCACCGCTGCTTGCCCACTGAATACATAATTTTTTCACTTGGAAATTGTTTTAGC Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
13 Some Problem Characteristics Genome sequence of 100Mb (C. elegans; yet relatively small) Can be interpreted in both directions The human genome is 35 larger Segment boundaries exhibit specific sequence patterns Almost every position is a potential segment start Many examples to classify Statistics within different segments differs Score segments of different length Segments are known to appear in a certain order Summary: BIG label sequence learning problem Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
14 Some Problem Characteristics Genome sequence of 100Mb (C. elegans; yet relatively small) Can be interpreted in both directions The human genome is 35 larger Segment boundaries exhibit specific sequence patterns Almost every position is a potential segment start Many examples to classify Statistics within different segments differs Score segments of different length Segments are known to appear in a certain order Summary: BIG label sequence learning problem Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
15 Some Problem Characteristics Genome sequence of 100Mb (C. elegans; yet relatively small) Can be interpreted in both directions The human genome is 35 larger Segment boundaries exhibit specific sequence patterns Almost every position is a potential segment start Many examples to classify Statistics within different segments differs Score segments of different length Segments are known to appear in a certain order Summary: BIG label sequence learning problem Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
16 Some Problem Characteristics Genome sequence of 100Mb (C. elegans; yet relatively small) Can be interpreted in both directions The human genome is 35 larger Segment boundaries exhibit specific sequence patterns Almost every position is a potential segment start Many examples to classify Statistics within different segments differs Score segments of different length Segments are known to appear in a certain order intergenic 5' UTR intron intron 3' UTR exon exon exon Summary: BIG label sequence learning problem intergenic Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
17 Some Problem Characteristics Genome sequence of 100Mb (C. elegans; yet relatively small) Can be interpreted in both directions The human genome is 35 larger Segment boundaries exhibit specific sequence patterns Almost every position is a potential segment start Many examples to classify Statistics within different segments differs Score segments of different length Segments are known to appear in a certain order 5' UTR intron intron 3' UTR intergenic exon exon exon intergenic Summary: BIG label sequence learning problem Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
18 Max-Margin Structured Output Learning Learn function f (y x) scoring segmentations y for input x Maximize f (y x) w.r.t. y for prediction: argmax f (y x) y Υ Idea: f (y x) f (ŷ x) for wrong labels ŷ y Approach: Given N sequence pairs (x 1, y 1 ),..., (x N, y N ) for training Solve using column-generation techniques: min f C N ξ n + P[f ] n=1 w.r.t. f (y n x n ) f (y x n ) l(y n, y) ξ n for all y n y Υ, n = 1,..., N All the remaining details are in f, P[f ], and l(, ). Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
19 Max-Margin Structured Output Learning Learn function f (y x) scoring segmentations y for input x Maximize f (y x) w.r.t. y for prediction: argmax f (y x) y Υ Idea: f (y x) f (ŷ x) for wrong labels ŷ y Approach: Given N sequence pairs (x 1, y 1 ),..., (x N, y N ) for training Solve using column-generation techniques: min f C N ξ n + P[f ] n=1 w.r.t. f (y n x n ) f (y x n ) l(y n, y) ξ n for all y n y Υ, n = 1,..., N All the remaining details are in f, P[f ], and l(, ). Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
20 Max-Margin Structured Output Learning Learn function f (y x) scoring segmentations y for input x Maximize f (y x) w.r.t. y for prediction: argmax f (y x) y Υ Idea: f (y x) f (ŷ x) for wrong labels ŷ y Approach: Given N sequence pairs (x 1, y 1 ),..., (x N, y N ) for training Solve using column-generation techniques: min f w.r.t. C N ξ n + P[f ] n=1 f (y n x n ) f (y x n ) l(y n, y) ξ n for all y n y Υ, n = 1,..., N All the remaining details are in f, P[f ], and l(, ). Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
21 Parametrization of f Requirements: Must allow efficient computation of argmax y Υ f (y x) Better has a small number of parameters Plausible model: Represent segmentation as sequence of segments: (p i, q i, y i ), for i = 1,..., I Model is additive in segment properties (Semi-Markov) f (y x) := I i=1 (f l y i (x pi ) + f s y i (x pi q i ) + f r y i (x qi )) Need to learn to score strings x! String kernels!?? No!... Yes! Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
22 Parametrization of f Requirements: Must allow efficient computation of argmax y Υ f (y x) Better has a small number of parameters Plausible model: Represent segmentation as sequence of segments: (p i, q i, y i ), for i = 1,..., I Model is additive in segment properties (Semi-Markov) f (y x) := I i=1 (f l y i (x pi ) + f s y i (x pi q i ) + f r y i (x qi )) Need to learn to score strings x! String kernels!?? No!... Yes! Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
23 Parametrization of f Requirements: Must allow efficient computation of argmax y Υ f (y x) Better has a small number of parameters Plausible model: Represent segmentation as sequence of segments: (p i, q i, y i ), for i = 1,..., I Model is additive in segment properties (Semi-Markov) f (y x) := I i=1 (f l y i (x pi ) + f s y i (x pi q i ) + f r y i (x qi )) Need to learn to score strings x! String kernels!?? No!... Yes! Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
24 Parametrization of f Requirements: Must allow efficient computation of argmax y Υ f (y x) Better has a small number of parameters Plausible model: Represent segmentation as sequence of segments: (p i, q i, y i ), for i = 1,..., I Model is additive in segment properties (Semi-Markov) f (y x) := I i=1 (f l y i (x pi ) + f s y i (x pi q i ) + f r y i (x qi )) Need to learn to score strings x! String kernels!?? No!... Yes! Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
25 Solve Problem in two Steps f (y x) := I i=1 (f l y i (x pi ) + f s y i (x pi q i ) + f r y i (x qi )) f y i (x ) := h y i (g y i (x )) Step 1: String analysis, leading to g yi : x R Step 2: Combination, leading to h yi : R R How to train g y i (x )? Should be large, if x is part of true label sequence Two-class problem: x at every possible position is negative, except at boundaries of true segments How to train h y i ( )? Simple 1-d function, e.g. piece-wise linear function Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
26 Solve Problem in two Steps f (y x) := I i=1 (f l y i (x pi ) + f s y i (x pi q i ) + f r y i (x qi )) f y i (x ) := h y i (g y i (x )) Step 1: String analysis, leading to g yi : x R Step 2: Combination, leading to h yi : R R How to train g y i (x )? Should be large, if x is part of true label sequence Two-class problem: x at every possible position is negative, except at boundaries of true segments How to train h y i ( )? Simple 1-d function, e.g. piece-wise linear function Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
27 Solve Problem in two Steps f (y x) := I i=1 (f l y i (x pi ) + f s y i (x pi q i ) + f r y i (x qi )) f y i (x ) := h y i (g y i (x )) Step 1: String analysis, leading to g yi : x R Step 2: Combination, leading to h yi : R R How to train g y i (x )? Should be large, if x is part of true label sequence Two-class problem: x at every possible position is negative, except at boundaries of true segments How to train h y i ( )? Simple 1-d function, e.g. piece-wise linear function Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
28 Discriminative Gene Prediction (simplified) [Rätsch, Sonnenburg, Srinivasan, Witte, Müller, Sommer, Schölkopf, 2007] Simplified Model: Score for splice form y = {(p j, q j )} J j=1 : J 1 F (y) := S GT (fj GT ) + j=1 J S AG (f AG j=2 j ) } {{ } Splice signals S LI (p j+1 q j ) + J 1 + j=1 J S LE (q j p j ) j=1 } {{ } Segment lengths Tune free parameters (in functions S GT, S AG, S LE, S LI ) by solving linear program using training set with known splice forms Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
29 Discriminative Gene Prediction (simplified) [Rätsch, Sonnenburg, Srinivasan, Witte, Müller, Sommer, Schölkopf, 2007] Simplified Model: Score for splice form y = {(p j, q j )} J j=1 : J 1 F (y) := S GT (fj GT ) + j=1 J S AG (f AG j=2 j ) } {{ } Splice signals S LI (p j+1 q j ) + J 1 + j=1 J S LE (q j p j ) j=1 } {{ } Segment lengths Tune free parameters (in functions S GT, S AG, S LE, S LI ) by solving linear program using training set with known splice forms Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
30 Example: Intron/Exon Boundary True Splice Sites CT...GTCGTA...GAAGCTAGGAGCGC...ACGCGT...GA 150 nucleotides window around dimer 1 GCCAATATTTTTCTATTCAGGTGCAATCAATCACCCATCAT 1 ATTGAATGAACATATTCCAGGGTCTCCTTCCACCTCAACAA 1 AGCAACGAACTCCATTACAGCAAGGACATCGAAGTCGATCA 1 GCCAATTTTTGACCTTGCAGAATCAATCGTGCACGTTCGGA -1 CATCTGAAATTTCCCCCAAGTATAGCGGAAATAGACCGACG -1 GAAATTTCCCCCAAGTATAGCGGAAATAGACCGACGAAATC -1 CCCAAGTATAGCGGAAATAGACCGACGAAATCGCTCTCTCC -1 AATCGCTCTCTCCCTGGGAGCGATGCGAATGTCAAATTCGA -1 ACCAAAAAATCAATTTTTAGATTTTTCGAATTAATTTTTCG -1 TGCTTTGCATGTTTCTAAAGTTACAGCCGTTCAAAATTTAA -1 GCATGTTTCTAAAGTTACAGCCGTTCAAAATTTAAAAACTC -1 ACCAATACGCAATGACTGAGTCTGTAATTTCACATAGTAAT Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
31 Gunnar Rätsch (FML, Tübingen) Large Scale Sequence. Analysis March 18, / 67 Example: Intron/Exon Boundary Potential Splice Sites CT...GTCGTA...GAAGCTAGGAGCGC...ACGCGT...GA 150 nucleotides window around dimer 1 GCCAATATTTTTCTATTCAGGTGCAATCAATCACCCATCAT 1 ATTGAATGAACATATTCCAGGGTCTCCTTCCACCTCAACAA 1 AGCAACGAACTCCATTACAGCAAGGACATCGAAGTCGATCA 1 GCCAATTTTTGACCTTGCAGAATCAATCGTGCACGTTCGGA -1 CATCTGAAATTTCCCCCAAGTATAGCGGAAATAGACCGACG -1 GAAATTTCCCCCAAGTATAGCGGAAATAGACCGACGAAATC -1 CCCAAGTATAGCGGAAATAGACCGACGAAATCGCTCTCTCC -1 AATCGCTCTCTCCCTGGGAGCGATGCGAATGTCAAATTCGA -1 ACCAAAAAATCAATTTTTAGATTTTTCGAATTAATTTTTCG -1 TGCTTTGCATGTTTCTAAAGTTACAGCCGTTCAAAATTTAA -1 GCATGTTTCTAAAGTTACAGCCGTTCAAAATTTAAAAACTC -1 ACCAATACGCAATGACTGAGTCTGTAATTTCACATAGTAAT
32 Example: Intron/Exon Boundary Potential Splice Sites CT...GTCGTA...GAAGCTAGGAGCGC...ACGCGT...GA 150 nucleotides window around dimer Basic idea: For instance, exploit: Exons have more G s and C s Certain motifs near boundary Sonnenburg, Schweikert et al Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
33 Substring Kernels General idea Count common substrings in two strings Sequences are deemed the more similar, the more common substrings they contain Variations Allow for gaps Include wildcards Allow for mismatches Include substitutions Motif Kernels Assign weights to substrings Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
34 Spectrum Kernel General idea [Leslie et al., 2002] For each k-mer s Σ k, the coordinate indexed by s will be the number of times s occurs in sequence x. Then the k-spectrum feature map is Φ Spectrum k (x) = (φ s (x)) s Σ k Here φ s (x) is the # occurrences of s in x. The spectrum kernel is now the inner product in the feature space defined by this map: k Spectrum (x, x ) = Φ Spectrum k (x), Φ Spectrum k (x ) Dimensionality: Exponential in k: Σ k Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
35 Simulation Example (Acceptor Splice Sites) Linear Kernel on GC-content features Spectrum kernel k Spectrum k (x, x ) Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
36 Position Dependence Given: Potential acceptor splice sites intron exon Goal: Rule that distinguishes true from false ones Position of motif is important ( T rich just before AG ) Spectrum kernel is blind w.r.t. positions New kernels for sequences with constant length Substring kernel per position (sum over positions) Oligo kernel Weighted Degree kernel Can detect motifs at specific positions weak if positions vary Extension: allow shifting Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
37 Weighted Degree Kernel [Rätsch and Sonnenburg, 2004] Equivalent to a mixture of spectrum kernels (up to order K) at every position for appropriately chosen β s: k(x i, x j ) = K k=1 L k+1 l=1 β k k Spectrum k (u l:l+k (x i ), u l:l+k (x j )) where β k = Pk K k+1 K k+1 = 2. (K k+1) k (k+1) Can be equivalently computed by k(x i, x j ) = K k=1 L k+1 l=1 β k I(u l:l+k (x i ) = u l:l+k (x j )) for appropriately chosen β k. Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
38 Weighted Degree Kernel Block Formulation Without shifts: Compare two sequences by identifying the largest matching blocks: where a matching block of length k implies many shorter matches: w k = min(k,k) j=1 β j (k j + 1). With shifts: Allows matching subsequences with offsets Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
39 Substring Kernel Comparison Linear kernel on GC-content features Spectrum kernel Weighted degree kernel Weighted degree kernel with shifts Remark: Higher order substring kernels typically exploit that correlations appear locally and not between arbitrary parts of the sequence (other than e.g. the polynomial kernel). Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
40 Fast string kernels? Use index structures to speed up computation Single kernel computation k(x, x ) = Φ(x), Φ(x ) Kernel (sub-)matrix k(x i, x j ), i I, j J Linear combination of kernel elements N N f (x) = α i k(x i, x) = α i Φ(x i ), Φ(x) i=1 Idea: Exploit that Φ(x) and also N i=1 α iφ(x i ) is sparse: Explicit maps Sorted lists (Suffix) trees/tries/arrays i=1 Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
41 Efficient data structures v = Φ(x) is very sparse Computation with v requires efficient operations on single dimensions, e.g. lookup v s or update v s = v s + α Use trees or arrays to store only non-zero elements Substring is the index into the tree or array Leads to more efficient optimization algorithms: Precompute v = N i=1 α iφ(x i ) Compute N i=1 α ik(x i, x) by s substring in x v s Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
42 Explicit Maps Require O( Σ k ) memory Explicitly store w = i α iφ(x i ) lookup and update operations are O(1) Updating all f (x i ) takes O(Q L k + N L k) Very efficient, but only work for small k Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
43 Sorted Lists Generate a sorted list with pairs (u, α) of length Q L O(Q L log(q L)) Requires O(Q L k) memory Iterate trough list and k-mer list of example (pre-sorted) identify co-occuring k-mers Single f (x i ) requires O((Q L log(q L) + L) k) All f (x i ) require O((Q L log(q L) + N L log(n L)) k) Requires additional sorting of long lists Also works for large k Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
44 Example: Trees & Tries Tree (trie) data structure stores sparse weightings on sequences (and their subsequences). Illustration: Three sequences AAA, AGA, GAA were added to a trie (α s are the weights of the sequences). Building tree: O(Q L k) Compute all f (x i ): O(N L k) Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
45 Solving the SVM Dual maximize N α i=1 α i 1 N N 2 i=1 j=1 α iα j y i y j k(x i, x j ) s.t. N i=1 α iy i = 0 0 α i C for i = 1, 2,..., N. Requires N 2 kernel computations expensive to compute (O(k L N 2 )) expensive to store matrix (O(N 2 )) Solving QP using interior point methods is expensive: O(N 3 ) Idea: Chunking based methods: Iterate Select small number of variables Optimize w.r.t. to these variables Stop if converged Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
46 Chunking F (α) := s.t. N i=1 α i 1 N N 2 i=1 j=1 α iα j y i y j k(x i, x j ) N i=1 α iy i = 0 0 α i C for i = 1, 2,..., N. Select Q variables i 1,..., i Q Random (inefficient) Sequential (inefficient) Heuristic selection motivated by KKT conditions Requires f (x j ) = N i=1 α ik(x i, x j ) for all j Points that have too small margin, but α i < C Points that are outside margin area, but α i > 0 Points with 0 α i C Solve QP of size Q (O(Q 3 )) Update f (x j ) if necessary Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
47 Chunking What do we need per iteration Compute f (x j ) = N i=1 α ik(x i, x j ) for all j Solve QP of size Q Complexity: O(Q N + Q 3 ) First part very expensive for large N Can we speedup computing f (x j )? So far for string kernels: O(Q N L k) With new data structures: O(N L k) Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
48 Algorithm INITIALIZATION f i = 0, α i = 0 for i = 1,..., N LOOP UNTIL CONVERGENCE For t = 1, 2,... Check optimality conditions and stop if optimal Select working set W based on g and α, store α old = α Solve reduced QP and update α clear w w w + (α j αj old )y j Φ(x j ) for all j W Update f i = f i + w, Φ(x i ) for all i = 1,..., N See Sonnenburg et al. [2007a] for more details. All implemented in Shogun toolbox ( Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
49 Human Splice Sites with WD Kernel Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
50 Example: Predictions in UCSC Browser Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
51 Example: Predictions in UCSC Browser Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
52 Integration of Signals DNA TSS Donor Acceptor Donor Acceptor polya/cleavage pre-mrna TIS Stop mrna cap polya Protein TSS TIS Stop cleave Don Acc Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
53 ngasp Competition Find the most accurate gene finder for annotation of new nematode genomes: Highly controlled competition conditions 4 Categories: Cat 1: Ab initio gene finders Cat 2: Dual/Multi-genome gene finders Cat 3: Gene finders that use EST/cDNA alignments Cat 4: Combining algorithms 47 submitted predictions from 17 different groups, including Fgenesh, Augustus, N-SCAN Evaluation on gold standard set of genes Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
54 Results: ngasp Nucleotide Exon Transcript Gene Cat. Method Avg Avg Avg Avg 1 mgene.init Craig EuGene Fgenesh Augustus mgene.multi N-SCAN EuGene mgene.seq Gramene Fgenesh Augustus mgene: Most accurate method in the ngasp genome annotation challenge for C. elegans Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
55 Results: mgene on Wormbase Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
56 Results: Genome-wide Predictions Annotation of other nematode genomes: (Schweikert et al., 2009) Genome Genome No. of No. exons/gene mgene best other size [Mbp] genes (mean) accuracy accuracy C. remanei % 93.8% C. japonica % 88.7% C. brenneri % 87.8% C. briggsae % 82.0% C.elegans model works well for closely related species. For intermediately distant organisms one can employ techniques to transfer learnt information. (Schweikert et al. 2009) For distantly related organisms retraining necessary: Galaxy based web service Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
57 Results: Genome-wide Predictions Annotation of other nematode genomes: (Schweikert et al., 2009) Genome Genome No. of No. exons/gene mgene best other size [Mbp] genes (mean) accuracy accuracy C. remanei % 93.8% C. japonica % 88.7% C. brenneri % 87.8% C. briggsae % 82.0% C.elegans model works well for closely related species. For intermediately distant organisms one can employ techniques to transfer learnt information. (Schweikert et al. 2009) For distantly related organisms retraining necessary: Galaxy based web service Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
58 Results: Genome-wide Predictions Annotation of other nematode genomes: (Schweikert et al., 2009) Genome Genome No. of No. exons/gene mgene best other size [Mbp] genes (mean) accuracy accuracy C. remanei % 93.8% C. japonica % 88.7% C. brenneri % 87.8% C. briggsae % 82.0% C.elegans model works well for closely related species. For intermediately distant organisms one can employ techniques to transfer learnt information. (Schweikert et al. 2009) For distantly related organisms retraining necessary: Galaxy based web service Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
59 mgene.web: Gene Finding for Everybody ;-) Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
60 mgene.web: Gene Finding for Everybody ;-) Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
61 Limitations/Extensions Gene finding accuracy still far from perfect Misses genes, predicts incorrect gene models Does not (yet) predict alternative transcripts Cannot predict when transcripts are expressed/modified/degraded... Accurate enough to accurately predict the effects of SNPs? Annotate all the newly sequenced variations of genomes Consensus site changes just the first step [Clark et al., 2007] Needs to be adapted to new genomes Requires sufficient number of known gene models for training Develop methods that exploit evolutionary information and gene models from other genomes [Schweikert et al., 2008] Model and understand the differences in transcription, RNA processing & translation. Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
62 Limitations/Extensions Gene finding accuracy still far from perfect Misses genes, predicts incorrect gene models Does not (yet) predict alternative transcripts Cannot predict when transcripts are expressed/modified/degraded... Accurate enough to accurately predict the effects of SNPs? Annotate all the newly sequenced variations of genomes Consensus site changes just the first step [Clark et al., 2007] Needs to be adapted to new genomes Requires sufficient number of known gene models for training Develop methods that exploit evolutionary information and gene models from other genomes [Schweikert et al., 2008] Model and understand the differences in transcription, RNA processing & translation. Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
63 Limitations/Extensions Gene finding accuracy still far from perfect Misses genes, predicts incorrect gene models Does not (yet) predict alternative transcripts Cannot predict when transcripts are expressed/modified/degraded... Accurate enough to accurately predict the effects of SNPs? Annotate all the newly sequenced variations of genomes Consensus site changes just the first step [Clark et al., 2007] Needs to be adapted to new genomes Requires sufficient number of known gene models for training Develop methods that exploit evolutionary information and gene models from other genomes [Schweikert et al., 2008] Model and understand the differences in transcription, RNA processing & translation. Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
64 Limitations/Extensions Gene finding accuracy still far from perfect Misses genes, predicts incorrect gene models Does not (yet) predict alternative transcripts Cannot predict when transcripts are expressed/modified/degraded... Accurate enough to accurately predict the effects of SNPs? Annotate all the newly sequenced variations of genomes Consensus site changes just the first step [Clark et al., 2007] Needs to be adapted to new genomes Requires sufficient number of known gene models for training Develop methods that exploit evolutionary information and gene models from other genomes [Schweikert et al., 2008] Model and understand the differences in transcription, RNA processing & translation. Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
65 Alternative Splicing: First Steps Predictions of alternative splicing Predict novel alternative splicing as independent events Use only information available to splicing machinery (Rätsch et. al, ISMB 05) Quite accurate for frequently appearing patterns Requires known gene structures Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
66 Alternative Splicing: More Steps Combine gene finding with prediction of single alternative splicing events Predict the splice graph of a gene Machine learning challenge: Input: DNA sequence Output: Splice graph msplicer approach can be extended to Include predictions of alternative splicing events Predict simple splice graphs Predicting arbitrary graphs is considerably harder Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
67 Domain Adaptation for Classification Motivation: Increasing number of sequenced genomes Often newly sequenced genomes are poorly annotated However often relatives with good annotation exist Idea: Transfer knowlege between organisms Study on domain adaptation for splice site prediction. Example: Splice site annotation in nematodes Newly sequenced organism: C. brennerei 590 confirmed splice site pairs Well annotated relative: C. elegans confirmed splice site pairs [Schweikert et al., NIPS 08] Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
68 Splice Site Recognition Idea: Discriminate true signal positions against all other positions Binary classification problem True sites: fixed window around a true splice site Decoy sites: all other consensus sites We learn a classification model from labeled training examples Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
69 Formal definition of Domain Adaptation Terminology: Well annotated organisms: Source domain Poorly annotated organisms: Target domain Distributional point of view: In Supervised Learning, example-label pairs are drawn from P(X, Y ) P S (X, Y ) might differ from P T (X, Y ) Factorization: P(X, Y ) = P(Y X ) P(X ) Covariate Shift: P S (X ) P T (X ) Differing Conditionals: P S (Y X ) P T (Y X ) Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
70 Splice Site Prediction and Domain Adaptation Sequence subject to opposing forces: P S (X ) P T (X ) Assume a splicesite pattern x occurs more frequently in a group of genes (e.g. chromosome) Duplication or deletion events could lead to altered P(X ) P S (Y X ) P T (Y X ) Think of the conditional as underlying mechanism Evolution of splicing machinery Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
71 Domain Adaptation Algorithms Overview Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
72 Domain Adaptation Methods Formula: 1 min w,b,ξ 2 wt w + C n i=1 s.t. y i (w T x i + b) + ξ i 1 0 i [1, n] ξ i ξ i 0 i [1, n] Resulting Model: f (x) = sign(w T x + b) Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
73 Domain Adaptation Methods Idea: Train on union of source and target Set trade-off via loss-term Formula: min w,b,ξ 1 2 wt w + C S n ξ i + C T i=1 m ξ i i=1 s.t. y i (w T x i + b) + ξ i 1 0 i [1, n + m] Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
74 Domain Adaptation Methods Idea: Combine trained models Efficient hyperparameter-optimization Formula: F (x) = αf S (x) + (1 α)f T (x) Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
75 Domain Adaptation Methods Idea: Takes interactions between source and target examples into account Two times linear search spaces for individual methods Captures General and Target-specific component Formula: F (x) = αf C (x) + (1 α)f T (x) Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
76 Domain Adaptation Methods Idea: Previous solution contains prior information Modified regularization term Formula: 1 min w T,ξ 2 wt T w T + C n ξ i Bw T T w S i=1 s.t. y i (w T T x i + b) + ξ i 1 0 i [1, n] Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
77 Domain Adaptation Methods Idea: Simultaneous optimization of both models Similarity between solution enforced Formula: min w S,w T,ξ m+n 1 2 w S w T 2 + C i=1 ξ i (1) s.t. y i ( w S, Φ(x i ) + b) 1 ξ i i 1,..., m y i ( w T, Φ(x i ) + b) 1 ξ i i m + 1,..., m + n Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
78 Domain Adaption Methods Idea: Match mean of source and target by reweighting examples Higher-order moments defined by mean when using a universal kernel [Huang, Smola, Gretton, Borgwardt, Schölkopf, 2007] Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
79 Domain Adaption Methods Idea: Based on same assumption Mean matching via translation rather than reweighting Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
80 Large scale experiments Varying distances Different data set sizes [MPI Developmental Biology, Departments 4 & 6 and UCSC Genome Browser] Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
81 Experimental Setup Source dataset size: always 100k examples Target dataset sizes: {2500, 6500, 16000, 64000, } Simple kernel (WDK of degree 1) Model selection for each method auroc/auprc measured for each setting Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
82 Results - Baseline Methods Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
83 Results - Improvement over Baseline Methods Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
84 Results - Summary Considerable improvements possible Sophisticated domain adaptation methods needed on distantly related organisms Best overall performance has DualTask Most cost effective Convex/AdvancedConvex Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
85 Domain Adaptation for LSL Problem: Very little data Relatively small amount of sequences available Only 50 well analysed genes Solution: Exploit that P. pacificus is closely related to C. elegans Can use C. elegans signal and content sensors But how can adapt the C. elegans parameters for gene structure prediction? Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
86 Domain Adaptation for LSL Problem: Very little data Relatively small amount of sequences available Only 50 well analysed genes Solution: Exploit that P. pacificus is closely related to C. elegans Can use C. elegans signal and content sensors But how can adapt the C. elegans parameters for gene structure prediction? Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
87 Domain Adaptation for LSL Problem: Very little data Relatively small amount of sequences available Only 50 well analysed genes Solution: Exploit that P. pacificus is closely related to C. elegans Can use C. elegans signal and content sensors But how can adapt the C. elegans parameters for gene structure prediction? Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
88 Domain Adaptation for LSL Problem: Very little data Relatively small amount of sequences available Only 50 well analysed genes Solution: Exploit that P. pacificus is closely related to C. elegans Can use C. elegans signal and content sensors But how can adapt the C. elegans parameters for gene structure prediction? Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
89 Domain Adaptation for LSL Details: Preliminary results! Signal predictions from C. elegans Training using 212 SNAP/EST gene models Parameter regularization against C. elegans solution Testing on 48 regions around known cdnas (±1000nt) GunnarSNAP Rätsch predictions (FML, Tübingen) provided by Christoph Dieterich Large Scale(Sommer Sequencelab) Analysis March 18, / 67
90 Domain Adaptation for LSL Details: Preliminary results! Signal predictions from C. elegans Training using 212 SNAP/EST gene models Parameter regularization against C. elegans solution Testing on 48 regions around known cdnas (±1000nt) GunnarSNAP Rätsch predictions (FML, Tübingen) provided by Christoph Dieterich Large Scale(Sommer Sequencelab) Analysis March 18, / 67
91 Summary and Future Work Genome Annotation is a huge structured output learning problem Proposed a two-step learning procedure separating the kernels from the structured output prediction Sequence classification already challenging (large!) String data structures make training feasible Gene prediction is more difficult in reality Predict splice graphs/alternative transcripts Regulation!? Auxiliary data!? Domain Adaptation First thorough comparison of Domain Adaptation Algorithms Learn models for multiple organisms simultaneously Develop more efficient training procedure Integrate these ideas into the gene finder Annotate the thousands of genomes that are currently being sequenced Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
92 Summary and Future Work Genome Annotation is a huge structured output learning problem Proposed a two-step learning procedure separating the kernels from the structured output prediction Sequence classification already challenging (large!) String data structures make training feasible Gene prediction is more difficult in reality Predict splice graphs/alternative transcripts Regulation!? Auxiliary data!? Domain Adaptation First thorough comparison of Domain Adaptation Algorithms Learn models for multiple organisms simultaneously Develop more efficient training procedure Integrate these ideas into the gene finder Annotate the thousands of genomes that are currently being sequenced Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
93 Summary and Future Work Genome Annotation is a huge structured output learning problem Proposed a two-step learning procedure separating the kernels from the structured output prediction Sequence classification already challenging (large!) String data structures make training feasible Gene prediction is more difficult in reality Predict splice graphs/alternative transcripts Regulation!? Auxiliary data!? Domain Adaptation First thorough comparison of Domain Adaptation Algorithms Learn models for multiple organisms simultaneously Develop more efficient training procedure Integrate these ideas into the gene finder Annotate the thousands of genomes that are currently being sequenced Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
94 Summary and Future Work Genome Annotation is a huge structured output learning problem Proposed a two-step learning procedure separating the kernels from the structured output prediction Sequence classification already challenging (large!) String data structures make training feasible Gene prediction is more difficult in reality Predict splice graphs/alternative transcripts Regulation!? Auxiliary data!? Domain Adaptation First thorough comparison of Domain Adaptation Algorithms Learn models for multiple organisms simultaneously Develop more efficient training procedure Integrate these ideas into the gene finder Annotate the thousands of genomes that are currently being sequenced Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
95 Summary and Future Work Genome Annotation is a huge structured output learning problem Proposed a two-step learning procedure separating the kernels from the structured output prediction Sequence classification already challenging (large!) String data structures make training feasible Gene prediction is more difficult in reality Predict splice graphs/alternative transcripts Regulation!? Auxiliary data!? Domain Adaptation First thorough comparison of Domain Adaptation Algorithms Learn models for multiple organisms simultaneously Develop more efficient training procedure Integrate these ideas into the gene finder Annotate the thousands of genomes that are currently being sequenced Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
96 Acknowledgments Sequence Analysis Sören Sonnenburg (FML/FIRST) Gabi Schweikert (FML/MPI) Alex Zien (FML & FIRST) Konrad Rieck (FIRST) Gene Finding Gabi Schweikert (FML/MPI) Jonas Behr (FML) Alex Zien (FML & FIRST) Georg Zeller (FML/MPI) Domain Adaptation Christian Widmer (FML) Gabi Schweikert (FML/MPI) Bernhard Schölkopf (MPI) More Information Slides with references are available online Thank you! Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
97 Acknowledgments Sequence Analysis Sören Sonnenburg (FML/FIRST) Gabi Schweikert (FML/MPI) Alex Zien (FML & FIRST) Konrad Rieck (FIRST) Gene Finding Gabi Schweikert (FML/MPI) Jonas Behr (FML) Alex Zien (FML & FIRST) Georg Zeller (FML/MPI) Domain Adaptation Christian Widmer (FML) Gabi Schweikert (FML/MPI) Bernhard Schölkopf (MPI) More Information Slides with references are available online Thank you! Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
98 Acknowledgments Sequence Analysis Sören Sonnenburg (FML/FIRST) Gabi Schweikert (FML/MPI) Alex Zien (FML & FIRST) Konrad Rieck (FIRST) Gene Finding Gabi Schweikert (FML/MPI) Jonas Behr (FML) Alex Zien (FML & FIRST) Georg Zeller (FML/MPI) Domain Adaptation Christian Widmer (FML) Gabi Schweikert (FML/MPI) Bernhard Schölkopf (MPI) More Information Slides with references are available online Thank you! Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
99 References I J. Behr, G. Schweikert, J. Cao, F. De Bona, G. Zeller, S. Laubinger, S. Ossowski, K. Schneeberger, D. Weigel, and G. Rätsch. Rna-seq and tiling arrays for improved gene finding. Presented at the CSHL Genome Informatics Meeting, May RM Clark, G Schweikert, C Toomajian, S Ossowski, G Zeller, P Shinn, N Warthmann, TT Hu, G Fu, DA Hinds, H Chen, KA Frazer, DH Huson, B Schölkopf, M Nordborg, G Rätsch, JR Ecker, and D Weigel. Common sequence polymorphisms shaping genetic diversity in arabidopsis thaliana. Science, 317(5836): , ISSN (Electronic). doi: /science C. Leslie, E. Eskin, and W. S. Noble. The spectrum kernel: A string kernel for SVM protein classification. In Proceedings of the Pacific Symposium on Biocomputing, pages , G. Rätsch and S. Sonnenburg. Accurate splice site detection for Caenorhabditis elegans. In K. Tsuda B. Schoelkopf and J.-P. Vert, editors, Kernel Methods in Computational Biology. MIT Press, G. Rätsch, S. Sonnenburg, and B. Schölkopf. RASE: recognition of alternatively spliced exons in C. elegans. Bioinformatics, 21(Suppl. 1):i369 i377, June G. Schweikert, G. Zeller, A. Zien, J. Behr, C.S. Ong, P. Philips, A. Bohlen, R. Bohnert, F. De Bona, S. Sonnenburg, and G. Rätsch. mgene: Accurate computational gene finding with application to nematode genomes. under revision, March Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
100 References II Gabriele Schweikert, Christian Widmer, Bernhard Schölkopf, and Gunnar Rätsch. An empirical analysis of domain adaptation algorithms. In Proc. NIPS 2008, Advances in Neural Information Processing Systems, accepted. S. Sonnenburg, G. Rätsch, A. Jagota, and K.-R. Müller. New methods for splice-site recognition. In Proc. International Conference on Artificial Neural Networks, S. Sonnenburg, G. Rätsch, and K. Rieck. Large Scale Kernel Machines, chapter Large Scale Learning with String Kernels. MIT Press, 2007a. S Sonnenburg, G Schweikert, P Philips, J Behr, and G Rätsch. Accurate splice site prediction using support vector machines. BMC Bioinformatics, 8 Suppl 10:S7, 2007b. ISSN (Electronic). doi: / S10-S7. Sören Sonnenburg, Alexander Zien, and Gunnar Rätsch. ARTS: Accurate recognition of transcription starts in human. Bioinformatics, 22(14):e , G Zeller, RM Clark, K Schneeberger, A Bohlen, D Weigel, and G Ratsch. Detecting polymorphic regions in arabidopsis thaliana with resequencing microarrays. Genome Res, 18 (6): , 2008a. ISSN (Print). doi: /gr G. Zeller, S.R. Henz, S. Laubinger, D. Weigel, and G Rätsch. Transcript normalization and segmentation of tiling array data. In Proceedings Pac. Symp. on Biocomputing, pages , 2008b. A. Zien, G. Rätsch, S. Mika, B. Schölkopf, T. Lengauer, and K.-R. Müller. Engineering Support Vector Machine Kernels That Recognize Translation Initiation Sites. BioInformatics, 16(9): , September Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
101 Results: RT-PCR Validation Validation of gene predictions for C. elegans: Schweikert et al., 2008 No. of genes No. of genes Frac. of genes analyzed w/ expression New genes 2, % Missing unconf. genes % new genes missed genes mgay_3 mgat_3 mgau_3 mgav_3 mgaw_3 mgax_3 mgaw_4 mgax_4 mgay_4 mgaz_4 mgbb_4 mgbd_4 Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
102 Domain Adaption by Learning vs. Homology Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
103 Domain Adaption by Learning vs. Homology Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
104 Domain Adaption by Learning vs. Homology Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
105 Domain Adaption by Learning vs. Homology Gunnar Rätsch (FML, Tübingen) Large Scale Sequence Analysis March 18, / 67
Large Scale Sequence Analysis with Applications to Genomics
 Large Scale Sequence Analysis with Applications to Genomics Gunnar Rätsch, Max Planck Society Tübingen, Germany Talk at CWI, Amsterdam October 23, 2009 Gunnar Rätsch (FML, Tübingen) Large Scale Sequence
Large Scale Sequence Analysis with Applications to Genomics Gunnar Rätsch, Max Planck Society Tübingen, Germany Talk at CWI, Amsterdam October 23, 2009 Gunnar Rätsch (FML, Tübingen) Large Scale Sequence
The CCR Model and Production Correspondence
 The CCR Model and Production Correspondence Tim Schöneberg The 19th of September Agenda Introduction Definitions Production Possiblity Set CCR Model and the Dual Problem Input excesses and output shortfalls
The CCR Model and Production Correspondence Tim Schöneberg The 19th of September Agenda Introduction Definitions Production Possiblity Set CCR Model and the Dual Problem Input excesses and output shortfalls
Efficiency change over time
 Efficiency change over time Heikki Tikanmäki Optimointiopin seminaari 14.11.2007 Contents Introduction (11.1) Window analysis (11.2) Example, application, analysis Malmquist index (11.3) Dealing with panel
Efficiency change over time Heikki Tikanmäki Optimointiopin seminaari 14.11.2007 Contents Introduction (11.1) Window analysis (11.2) Example, application, analysis Malmquist index (11.3) Dealing with panel
Returns to Scale II. S ysteemianalyysin. Laboratorio. Esitelmä 8 Timo Salminen. Teknillinen korkeakoulu
 Returns to Scale II Contents Most Productive Scale Size Further Considerations Relaxation of the Convexity Condition Useful Reminder Theorem 5.5 A DMU found to be efficient with a CCR model will also be
Returns to Scale II Contents Most Productive Scale Size Further Considerations Relaxation of the Convexity Condition Useful Reminder Theorem 5.5 A DMU found to be efficient with a CCR model will also be
Alternative DEA Models
 Mat-2.4142 Alternative DEA Models 19.9.2007 Table of Contents Banker-Charnes-Cooper Model Additive Model Example Data Home assignment BCC Model (Banker-Charnes-Cooper) production frontiers spanned by convex
Mat-2.4142 Alternative DEA Models 19.9.2007 Table of Contents Banker-Charnes-Cooper Model Additive Model Example Data Home assignment BCC Model (Banker-Charnes-Cooper) production frontiers spanned by convex
Capacity Utilization
 Capacity Utilization Tim Schöneberg 28th November Agenda Introduction Fixed and variable input ressources Technical capacity utilization Price based capacity utilization measure Long run and short run
Capacity Utilization Tim Schöneberg 28th November Agenda Introduction Fixed and variable input ressources Technical capacity utilization Price based capacity utilization measure Long run and short run
Other approaches to restrict multipliers
 Other approaches to restrict multipliers Heikki Tikanmäki Optimointiopin seminaari 10.10.2007 Contents Short revision (6.2) Another Assurance Region Model (6.3) Cone-Ratio Method (6.4) An Application of
Other approaches to restrict multipliers Heikki Tikanmäki Optimointiopin seminaari 10.10.2007 Contents Short revision (6.2) Another Assurance Region Model (6.3) Cone-Ratio Method (6.4) An Application of
Bounds on non-surjective cellular automata
 Bounds on non-surjective cellular automata Jarkko Kari Pascal Vanier Thomas Zeume University of Turku LIF Marseille Universität Hannover 27 august 2009 J. Kari, P. Vanier, T. Zeume (UTU) Bounds on non-surjective
Bounds on non-surjective cellular automata Jarkko Kari Pascal Vanier Thomas Zeume University of Turku LIF Marseille Universität Hannover 27 august 2009 J. Kari, P. Vanier, T. Zeume (UTU) Bounds on non-surjective
Information on preparing Presentation
 Information on preparing Presentation Seminar on big data management Lecturer: Spring 2017 20.1.2017 1 Agenda Hints and tips on giving a good presentation Watch two videos and discussion 22.1.2017 2 Goals
Information on preparing Presentation Seminar on big data management Lecturer: Spring 2017 20.1.2017 1 Agenda Hints and tips on giving a good presentation Watch two videos and discussion 22.1.2017 2 Goals
Statistical design. Tuomas Selander
 Statistical design Tuomas Selander 28.8.2014 Introduction Biostatistician Work area KYS-erva KYS, Jyväskylä, Joensuu, Mikkeli, Savonlinna Work tasks Statistical methods, selection and quiding Data analysis
Statistical design Tuomas Selander 28.8.2014 Introduction Biostatistician Work area KYS-erva KYS, Jyväskylä, Joensuu, Mikkeli, Savonlinna Work tasks Statistical methods, selection and quiding Data analysis
Gap-filling methods for CH 4 data
 Gap-filling methods for CH 4 data Sigrid Dengel University of Helsinki Outline - Ecosystems known for CH 4 emissions; - Why is gap-filling of CH 4 data not as easy and straight forward as CO 2 ; - Gap-filling
Gap-filling methods for CH 4 data Sigrid Dengel University of Helsinki Outline - Ecosystems known for CH 4 emissions; - Why is gap-filling of CH 4 data not as easy and straight forward as CO 2 ; - Gap-filling
On instrument costs in decentralized macroeconomic decision making (Helsingin Kauppakorkeakoulun julkaisuja ; D-31)
 On instrument costs in decentralized macroeconomic decision making (Helsingin Kauppakorkeakoulun julkaisuja ; D-31) Juha Kahkonen Click here if your download doesn"t start automatically On instrument costs
On instrument costs in decentralized macroeconomic decision making (Helsingin Kauppakorkeakoulun julkaisuja ; D-31) Juha Kahkonen Click here if your download doesn"t start automatically On instrument costs
Functional Genomics & Proteomics
 Functional Genomics & Proteomics Genome Sequences TCACAATTTAGACATCTAGTCTTCCACTTAAGCATATTTAGATTGTTTCCAGTTTTCAGCTTTTATGACTAAATCTTCTAAAATTGTTTTTCCCTAAATGTATATTTTAATTTGTCTCAGGAGTAGAATTTCTGAGTCATAAAGCGGT CATATGTATAAATTTTAGGTGCCTCATAGCTCTTCAAATAGTCATCCCATTTTATACATCCAGGCAATATATGAGAGTTCTTGGTGCTCCACATCTTAGCTAGGATTTGATGTCAACCAGTCTCTTTAATTTAGATATTCTAGTACAT
Functional Genomics & Proteomics Genome Sequences TCACAATTTAGACATCTAGTCTTCCACTTAAGCATATTTAGATTGTTTCCAGTTTTCAGCTTTTATGACTAAATCTTCTAAAATTGTTTTTCCCTAAATGTATATTTTAATTTGTCTCAGGAGTAGAATTTCTGAGTCATAAAGCGGT CATATGTATAAATTTTAGGTGCCTCATAGCTCTTCAAATAGTCATCCCATTTTATACATCCAGGCAATATATGAGAGTTCTTGGTGCTCCACATCTTAGCTAGGATTTGATGTCAACCAGTCTCTTTAATTTAGATATTCTAGTACAT
7.4 Variability management
 7.4 Variability management time... space software product-line should support variability in space (different products) support variability in time (maintenance, evolution) 1 Product variation Product
7.4 Variability management time... space software product-line should support variability in space (different products) support variability in time (maintenance, evolution) 1 Product variation Product
16. Allocation Models
 16. Allocation Models Juha Saloheimo 17.1.27 S steemianalsin Optimointiopin seminaari - Sks 27 Content Introduction Overall Efficienc with common prices and costs Cost Efficienc S steemianalsin Revenue
16. Allocation Models Juha Saloheimo 17.1.27 S steemianalsin Optimointiopin seminaari - Sks 27 Content Introduction Overall Efficienc with common prices and costs Cost Efficienc S steemianalsin Revenue
Genome 373: Genomic Informatics. Professors Elhanan Borenstein and Jay Shendure
 Genome 373: Genomic Informatics Professors Elhanan Borenstein and Jay Shendure Genome 373 This course is intended to introduce students to the breadth of problems and methods in computational analysis
Genome 373: Genomic Informatics Professors Elhanan Borenstein and Jay Shendure Genome 373 This course is intended to introduce students to the breadth of problems and methods in computational analysis
Chapter 7. Motif finding (week 11) Chapter 8. Sequence binning (week 11)
 Course organization Introduction ( Week 1) Part I: Algorithms for Sequence Analysis (Week 1-11) Chapter 1-3, Models and theories» Probability theory and Statistics (Week 2)» Algorithm complexity analysis
Course organization Introduction ( Week 1) Part I: Algorithms for Sequence Analysis (Week 1-11) Chapter 1-3, Models and theories» Probability theory and Statistics (Week 2)» Algorithm complexity analysis
Results on the new polydrug use questions in the Finnish TDI data
 Results on the new polydrug use questions in the Finnish TDI data Multi-drug use, polydrug use and problematic polydrug use Martta Forsell, Finnish Focal Point 28/09/2015 Martta Forsell 1 28/09/2015 Esityksen
Results on the new polydrug use questions in the Finnish TDI data Multi-drug use, polydrug use and problematic polydrug use Martta Forsell, Finnish Focal Point 28/09/2015 Martta Forsell 1 28/09/2015 Esityksen
Basset: Learning the regulatory code of the accessible genome with deep convolutional neural networks. David R. Kelley
 Basset: Learning the regulatory code of the accessible genome with deep convolutional neural networks David R. Kelley DNA codes for complex life. How? Kundaje et al. Integrative analysis of 111 reference
Basset: Learning the regulatory code of the accessible genome with deep convolutional neural networks David R. Kelley DNA codes for complex life. How? Kundaje et al. Integrative analysis of 111 reference
The Viking Battle - Part Version: Finnish
 The Viking Battle - Part 1 015 Version: Finnish Tehtävä 1 Olkoon kokonaisluku, ja olkoon A n joukko A n = { n k k Z, 0 k < n}. Selvitä suurin kokonaisluku M n, jota ei voi kirjoittaa yhden tai useamman
The Viking Battle - Part 1 015 Version: Finnish Tehtävä 1 Olkoon kokonaisluku, ja olkoon A n joukko A n = { n k k Z, 0 k < n}. Selvitä suurin kokonaisluku M n, jota ei voi kirjoittaa yhden tai useamman
Alternatives to the DFT
 Alternatives to the DFT Doru Balcan Carnegie Mellon University joint work with Aliaksei Sandryhaila, Jonathan Gross, and Markus Püschel - appeared in IEEE ICASSP 08 - Introduction Discrete time signal
Alternatives to the DFT Doru Balcan Carnegie Mellon University joint work with Aliaksei Sandryhaila, Jonathan Gross, and Markus Püschel - appeared in IEEE ICASSP 08 - Introduction Discrete time signal
Plasmid Name: pmm290. Aliases: none known. Length: bp. Constructed by: Mike Moser/Cristina Swanson. Last updated: 17 August 2009
 Plasmid Name: pmm290 Aliases: none known Length: 11707 bp Constructed by: Mike Moser/Cristina Swanson Last updated: 17 August 2009 Description and application: This is a mammalian expression vector for
Plasmid Name: pmm290 Aliases: none known Length: 11707 bp Constructed by: Mike Moser/Cristina Swanson Last updated: 17 August 2009 Description and application: This is a mammalian expression vector for
Capacity utilization
 Mat-2.4142 Seminar on optimization Capacity utilization 12.12.2007 Contents Summary of chapter 14 Related DEA-solver models Illustrative examples Measure of technical capacity utilization Price-based measure
Mat-2.4142 Seminar on optimization Capacity utilization 12.12.2007 Contents Summary of chapter 14 Related DEA-solver models Illustrative examples Measure of technical capacity utilization Price-based measure
ECVETin soveltuvuus suomalaisiin tutkinnon perusteisiin. Case:Yrittäjyyskurssi matkailualan opiskelijoille englantilaisen opettajan toteuttamana
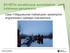 ECVETin soveltuvuus suomalaisiin tutkinnon perusteisiin Case:Yrittäjyyskurssi matkailualan opiskelijoille englantilaisen opettajan toteuttamana Taustaa KAO mukana FINECVET-hankeessa, jossa pilotoimme ECVETiä
ECVETin soveltuvuus suomalaisiin tutkinnon perusteisiin Case:Yrittäjyyskurssi matkailualan opiskelijoille englantilaisen opettajan toteuttamana Taustaa KAO mukana FINECVET-hankeessa, jossa pilotoimme ECVETiä
Searching (Sub-)Strings. Ulf Leser
 Searching (Sub-)Strings Ulf Leser This Lecture Exact substring search Naïve Boyer-Moore Searching with profiles Sequence profiles Ungapped approximate search Statistical evaluation of search results Ulf
Searching (Sub-)Strings Ulf Leser This Lecture Exact substring search Naïve Boyer-Moore Searching with profiles Sequence profiles Ungapped approximate search Statistical evaluation of search results Ulf
SIMULINK S-funktiot. SIMULINK S-funktiot
 S-funktio on ohjelmointikielellä (Matlab, C, Fortran) laadittu oma algoritmi tai dynaamisen järjestelmän kuvaus, jota voidaan käyttää Simulink-malleissa kuin mitä tahansa valmista lohkoa. S-funktion rakenne
S-funktio on ohjelmointikielellä (Matlab, C, Fortran) laadittu oma algoritmi tai dynaamisen järjestelmän kuvaus, jota voidaan käyttää Simulink-malleissa kuin mitä tahansa valmista lohkoa. S-funktion rakenne
On instrument costs in decentralized macroeconomic decision making (Helsingin Kauppakorkeakoulun julkaisuja ; D-31)
 On instrument costs in decentralized macroeconomic decision making (Helsingin Kauppakorkeakoulun julkaisuja ; D-31) Juha Kahkonen Click here if your download doesn"t start automatically On instrument costs
On instrument costs in decentralized macroeconomic decision making (Helsingin Kauppakorkeakoulun julkaisuja ; D-31) Juha Kahkonen Click here if your download doesn"t start automatically On instrument costs
1.3Lohkorakenne muodostetaan käyttämällä a) puolipistettä b) aaltosulkeita c) BEGIN ja END lausekkeita d) sisennystä
 OULUN YLIOPISTO Tietojenkäsittelytieteiden laitos Johdatus ohjelmointiin 81122P (4 ov.) 30.5.2005 Ohjelmointikieli on Java. Tentissä saa olla materiaali mukana. Tenttitulokset julkaistaan aikaisintaan
OULUN YLIOPISTO Tietojenkäsittelytieteiden laitos Johdatus ohjelmointiin 81122P (4 ov.) 30.5.2005 Ohjelmointikieli on Java. Tentissä saa olla materiaali mukana. Tenttitulokset julkaistaan aikaisintaan
Valuation of Asian Quanto- Basket Options
 Valuation of Asian Quanto- Basket Options (Final Presentation) 21.11.2011 Thesis Instructor and Supervisor: Prof. Ahti Salo Työn saa tallentaa ja julkistaa Aalto-yliopiston avoimilla verkkosivuilla. Muilta
Valuation of Asian Quanto- Basket Options (Final Presentation) 21.11.2011 Thesis Instructor and Supervisor: Prof. Ahti Salo Työn saa tallentaa ja julkistaa Aalto-yliopiston avoimilla verkkosivuilla. Muilta
Uusi Ajatus Löytyy Luonnosta 4 (käsikirja) (Finnish Edition)
 Uusi Ajatus Löytyy Luonnosta 4 (käsikirja) (Finnish Edition) Esko Jalkanen Click here if your download doesn"t start automatically Uusi Ajatus Löytyy Luonnosta 4 (käsikirja) (Finnish Edition) Esko Jalkanen
Uusi Ajatus Löytyy Luonnosta 4 (käsikirja) (Finnish Edition) Esko Jalkanen Click here if your download doesn"t start automatically Uusi Ajatus Löytyy Luonnosta 4 (käsikirja) (Finnish Edition) Esko Jalkanen
Operatioanalyysi 2011, Harjoitus 4, viikko 40
 Operatioanalyysi 2011, Harjoitus 4, viikko 40 H4t1, Exercise 4.2. H4t2, Exercise 4.3. H4t3, Exercise 4.4. H4t4, Exercise 4.5. H4t5, Exercise 4.6. (Exercise 4.2.) 1 4.2. Solve the LP max z = x 1 + 2x 2
Operatioanalyysi 2011, Harjoitus 4, viikko 40 H4t1, Exercise 4.2. H4t2, Exercise 4.3. H4t3, Exercise 4.4. H4t4, Exercise 4.5. H4t5, Exercise 4.6. (Exercise 4.2.) 1 4.2. Solve the LP max z = x 1 + 2x 2
1. SIT. The handler and dog stop with the dog sitting at heel. When the dog is sitting, the handler cues the dog to heel forward.
 START START SIT 1. SIT. The handler and dog stop with the dog sitting at heel. When the dog is sitting, the handler cues the dog to heel forward. This is a static exercise. SIT STAND 2. SIT STAND. The
START START SIT 1. SIT. The handler and dog stop with the dog sitting at heel. When the dog is sitting, the handler cues the dog to heel forward. This is a static exercise. SIT STAND 2. SIT STAND. The
Categorical Decision Making Units and Comparison of Efficiency between Different Systems
 Categorical Decision Making Units and Comparison of Efficiency between Different Systems Mat-2.4142 Optimointiopin Seminaari Source William W. Cooper, Lawrence M. Seiford, Kaoru Tone: Data Envelopment
Categorical Decision Making Units and Comparison of Efficiency between Different Systems Mat-2.4142 Optimointiopin Seminaari Source William W. Cooper, Lawrence M. Seiford, Kaoru Tone: Data Envelopment
On instrument costs in decentralized macroeconomic decision making (Helsingin Kauppakorkeakoulun julkaisuja ; D-31)
 On instrument costs in decentralized macroeconomic decision making (Helsingin Kauppakorkeakoulun julkaisuja ; D-31) Juha Kahkonen Click here if your download doesn"t start automatically On instrument costs
On instrument costs in decentralized macroeconomic decision making (Helsingin Kauppakorkeakoulun julkaisuja ; D-31) Juha Kahkonen Click here if your download doesn"t start automatically On instrument costs
anna minun kertoa let me tell you
 anna minun kertoa let me tell you anna minun kertoa I OSA 1. Anna minun kertoa sinulle mitä oli. Tiedän että osaan. Kykenen siihen. Teen nyt niin. Minulla on oikeus. Sanani voivat olla puutteellisia mutta
anna minun kertoa let me tell you anna minun kertoa I OSA 1. Anna minun kertoa sinulle mitä oli. Tiedän että osaan. Kykenen siihen. Teen nyt niin. Minulla on oikeus. Sanani voivat olla puutteellisia mutta
LX 70. Ominaisuuksien mittaustulokset 1-kerroksinen 2-kerroksinen. Fyysiset ominaisuudet, nimellisarvot. Kalvon ominaisuudet
 LX 70 % Läpäisy 36 32 % Absorptio 30 40 % Heijastus 34 28 % Läpäisy 72 65 % Heijastus ulkopuoli 9 16 % Heijastus sisäpuoli 9 13 Emissiivisyys.77.77 Auringonsuojakerroin.54.58 Auringonsäteilyn lämmönsiirtokerroin.47.50
LX 70 % Läpäisy 36 32 % Absorptio 30 40 % Heijastus 34 28 % Läpäisy 72 65 % Heijastus ulkopuoli 9 16 % Heijastus sisäpuoli 9 13 Emissiivisyys.77.77 Auringonsuojakerroin.54.58 Auringonsäteilyn lämmönsiirtokerroin.47.50
Information on Finnish Language Courses Spring Semester 2018 Päivi Paukku & Jenni Laine Centre for Language and Communication Studies
 Information on Finnish Language Courses Spring Semester 2018 Päivi Paukku & Jenni Laine 4.1.2018 Centre for Language and Communication Studies Puhutko suomea? -Hei! -Hei hei! -Moi! -Moi moi! -Terve! -Terve
Information on Finnish Language Courses Spring Semester 2018 Päivi Paukku & Jenni Laine 4.1.2018 Centre for Language and Communication Studies Puhutko suomea? -Hei! -Hei hei! -Moi! -Moi moi! -Terve! -Terve
State of the Union... Functional Genomics Research Stream. Molecular Biology. Genomics. Computational Biology
 Functional Genomics Research Stream State of the Union... Research Meeting: February 16, 2010 Functional Genomics & Research Report III Concepts Genomics Molecular Biology Computational Biology Genome
Functional Genomics Research Stream State of the Union... Research Meeting: February 16, 2010 Functional Genomics & Research Report III Concepts Genomics Molecular Biology Computational Biology Genome
Mat Seminar on Optimization. Data Envelopment Analysis. Economies of Scope S ysteemianalyysin. Laboratorio. Teknillinen korkeakoulu
 Mat-2.4142 Seminar on Optimization Data Envelopment Analysis Economies of Scope 21.11.2007 Economies of Scope Introduced 1982 by Panzar and Willing Support decisions like: Should a firm... Produce a variety
Mat-2.4142 Seminar on Optimization Data Envelopment Analysis Economies of Scope 21.11.2007 Economies of Scope Introduced 1982 by Panzar and Willing Support decisions like: Should a firm... Produce a variety
Choose Finland-Helsinki Valitse Finland-Helsinki
 Write down the Temporary Application ID. If you do not manage to complete the form you can continue where you stopped with this ID no. Muista Temporary Application ID. Jos et onnistu täyttää lomake loppuun
Write down the Temporary Application ID. If you do not manage to complete the form you can continue where you stopped with this ID no. Muista Temporary Application ID. Jos et onnistu täyttää lomake loppuun
Rekisteröiminen - FAQ
 Rekisteröiminen - FAQ Miten Akun/laturin rekisteröiminen tehdään Akun/laturin rekisteröiminen tapahtuu samalla tavalla kuin nykyinen takuurekisteröityminen koneille. Nykyistä tietokantaa on muokattu niin,
Rekisteröiminen - FAQ Miten Akun/laturin rekisteröiminen tehdään Akun/laturin rekisteröiminen tapahtuu samalla tavalla kuin nykyinen takuurekisteröityminen koneille. Nykyistä tietokantaa on muokattu niin,
Network to Get Work. Tehtäviä opiskelijoille Assignments for students. www.laurea.fi
 Network to Get Work Tehtäviä opiskelijoille Assignments for students www.laurea.fi Ohje henkilöstölle Instructions for Staff Seuraavassa on esitetty joukko tehtäviä, joista voit valita opiskelijaryhmällesi
Network to Get Work Tehtäviä opiskelijoille Assignments for students www.laurea.fi Ohje henkilöstölle Instructions for Staff Seuraavassa on esitetty joukko tehtäviä, joista voit valita opiskelijaryhmällesi
LYTH-CONS CONSISTENCY TRANSMITTER
 LYTH-CONS CONSISTENCY TRANSMITTER LYTH-INSTRUMENT OY has generate new consistency transmitter with blade-system to meet high technical requirements in Pulp&Paper industries. Insurmountable advantages are
LYTH-CONS CONSISTENCY TRANSMITTER LYTH-INSTRUMENT OY has generate new consistency transmitter with blade-system to meet high technical requirements in Pulp&Paper industries. Insurmountable advantages are
Information on Finnish Language Courses Spring Semester 2017 Jenni Laine
 Information on Finnish Language Courses Spring Semester 2017 Jenni Laine 4.1.2017 KIELIKESKUS LANGUAGE CENTRE Puhutko suomea? Do you speak Finnish? -Hei! -Moi! -Mitä kuuluu? -Kiitos, hyvää. -Entä sinulle?
Information on Finnish Language Courses Spring Semester 2017 Jenni Laine 4.1.2017 KIELIKESKUS LANGUAGE CENTRE Puhutko suomea? Do you speak Finnish? -Hei! -Moi! -Mitä kuuluu? -Kiitos, hyvää. -Entä sinulle?
Strict singularity of a Volterra-type integral operator on H p
 Strict singularity of a Volterra-type integral operator on H p Santeri Miihkinen, University of Helsinki IWOTA St. Louis, 18-22 July 2016 Santeri Miihkinen, University of Helsinki Volterra-type integral
Strict singularity of a Volterra-type integral operator on H p Santeri Miihkinen, University of Helsinki IWOTA St. Louis, 18-22 July 2016 Santeri Miihkinen, University of Helsinki Volterra-type integral
FETAL FIBROBLASTS, PASSAGE 10
 Double-stranded methylation patterns of a 104-bp L1 promoter in DNAs from fetal fibroblast passages 10, 14, 17, and 22 using barcoded hairpinbisulfite PCR. Fifteen L1 sequences were analyzed for passages
Double-stranded methylation patterns of a 104-bp L1 promoter in DNAs from fetal fibroblast passages 10, 14, 17, and 22 using barcoded hairpinbisulfite PCR. Fifteen L1 sequences were analyzed for passages
C++11 seminaari, kevät Johannes Koskinen
 C++11 seminaari, kevät 2012 Johannes Koskinen Sisältö Mikä onkaan ongelma? Standardidraftin luku 29: Atomiset tyypit Muistimalli Rinnakkaisuus On multicore systems, when a thread writes a value to memory,
C++11 seminaari, kevät 2012 Johannes Koskinen Sisältö Mikä onkaan ongelma? Standardidraftin luku 29: Atomiset tyypit Muistimalli Rinnakkaisuus On multicore systems, when a thread writes a value to memory,
7. Product-line architectures
 7. Product-line architectures 7.1 Introduction 7.2 Product-line basics 7.3 Layered style for product-lines 7.4 Variability management 7.5 Benefits and problems with product-lines 1 Short history of software
7. Product-line architectures 7.1 Introduction 7.2 Product-line basics 7.3 Layered style for product-lines 7.4 Variability management 7.5 Benefits and problems with product-lines 1 Short history of software
Tilausvahvistus. Anttolan Urheilijat HENNA-RIIKKA HAIKONEN KUMMANNIEMENTIE 5 B RAHULA. Anttolan Urheilijat
 7.80.4 Asiakasnumero: 3000359 KALLE MANNINEN KOVASTENLUODONTIE 46 51600 HAUKIVUORI Toimitusosoite: KUMMANNIEMENTIE 5 B 51720 RAHULA Viitteenne: Henna-Riikka Haikonen Viitteemme: Pyry Niemi +358400874498
7.80.4 Asiakasnumero: 3000359 KALLE MANNINEN KOVASTENLUODONTIE 46 51600 HAUKIVUORI Toimitusosoite: KUMMANNIEMENTIE 5 B 51720 RAHULA Viitteenne: Henna-Riikka Haikonen Viitteemme: Pyry Niemi +358400874498
MALE ADULT FIBROBLAST LINE (82-6hTERT)
 Double-stranded methylation patterns of a 104-bp L1 promoter in DNAs from male and female fibroblasts, male leukocytes and female lymphoblastoid cells using hairpin-bisulfite PCR. Fifteen L1 sequences
Double-stranded methylation patterns of a 104-bp L1 promoter in DNAs from male and female fibroblasts, male leukocytes and female lymphoblastoid cells using hairpin-bisulfite PCR. Fifteen L1 sequences
812336A C++ -kielen perusteet, 21.8.2010
 812336A C++ -kielen perusteet, 21.8.2010 1. Vastaa lyhyesti seuraaviin kysymyksiin (1p kaikista): a) Mitä tarkoittaa funktion ylikuormittaminen (overloading)? b) Mitä tarkoittaa jäsenfunktion ylimääritys
812336A C++ -kielen perusteet, 21.8.2010 1. Vastaa lyhyesti seuraaviin kysymyksiin (1p kaikista): a) Mitä tarkoittaa funktion ylikuormittaminen (overloading)? b) Mitä tarkoittaa jäsenfunktion ylimääritys
Toppila/Kivistö 10.01.2013 Vastaa kaikkin neljään tehtävään, jotka kukin arvostellaan asteikolla 0-6 pistettä.
 ..23 Vastaa kaikkin neljään tehtävään, jotka kukin arvostellaan asteikolla -6 pistettä. Tehtävä Ovatko seuraavat väittämät oikein vai väärin? Perustele vastauksesi. (a) Lineaarisen kokonaislukutehtävän
..23 Vastaa kaikkin neljään tehtävään, jotka kukin arvostellaan asteikolla -6 pistettä. Tehtävä Ovatko seuraavat väittämät oikein vai väärin? Perustele vastauksesi. (a) Lineaarisen kokonaislukutehtävän
Information on Finnish Courses Autumn Semester 2017 Jenni Laine & Päivi Paukku Centre for Language and Communication Studies
 Information on Finnish Courses Autumn Semester 2017 Jenni Laine & Päivi Paukku 24.8.2017 Centre for Language and Communication Studies Puhutko suomea? -Hei! -Hei hei! -Moi! -Moi moi! -Terve! -Terve terve!
Information on Finnish Courses Autumn Semester 2017 Jenni Laine & Päivi Paukku 24.8.2017 Centre for Language and Communication Studies Puhutko suomea? -Hei! -Hei hei! -Moi! -Moi moi! -Terve! -Terve terve!
Constructive Alignment in Specialisation Studies in Industrial Pharmacy in Finland
 Constructive Alignment in Specialisation Studies in Industrial Pharmacy in Finland Anne Mari Juppo, Nina Katajavuori University of Helsinki Faculty of Pharmacy 23.7.2012 1 Background Pedagogic research
Constructive Alignment in Specialisation Studies in Industrial Pharmacy in Finland Anne Mari Juppo, Nina Katajavuori University of Helsinki Faculty of Pharmacy 23.7.2012 1 Background Pedagogic research
Tietorakenteet ja algoritmit
 Tietorakenteet ja algoritmit Taulukon edut Taulukon haitat Taulukon haittojen välttäminen Dynaamisesti linkattu lista Linkatun listan solmun määrittelytavat Lineaarisen listan toteutus dynaamisesti linkattuna
Tietorakenteet ja algoritmit Taulukon edut Taulukon haitat Taulukon haittojen välttäminen Dynaamisesti linkattu lista Linkatun listan solmun määrittelytavat Lineaarisen listan toteutus dynaamisesti linkattuna
Land-Use Model for the Helsinki Metropolitan Area
 Land-Use Model for the Helsinki Metropolitan Area Paavo Moilanen Introduction & Background Metropolitan Area Council asked 2005: What is good land use for the transport systems plan? At first a literature
Land-Use Model for the Helsinki Metropolitan Area Paavo Moilanen Introduction & Background Metropolitan Area Council asked 2005: What is good land use for the transport systems plan? At first a literature
FIS IMATRAN KYLPYLÄHIIHDOT Team captains meeting
 FIS IMATRAN KYLPYLÄHIIHDOT 8.-9.12.2018 Team captains meeting 8.12.2018 Agenda 1 Opening of the meeting 2 Presence 3 Organizer s personell 4 Jury 5 Weather forecast 6 Composition of competitors startlists
FIS IMATRAN KYLPYLÄHIIHDOT 8.-9.12.2018 Team captains meeting 8.12.2018 Agenda 1 Opening of the meeting 2 Presence 3 Organizer s personell 4 Jury 5 Weather forecast 6 Composition of competitors startlists
11/17/11. Gene Regulation. Gene Regulation. Gene Regulation. Finding Regulatory Motifs in DNA Sequences. Regulatory Proteins
 Gene Regulation Finding Regulatory Motifs in DNA Sequences An experiment shows that when X is knocked out, 20 other s are not expressed How can one have such drastic effects? Regulatory Proteins Gene X
Gene Regulation Finding Regulatory Motifs in DNA Sequences An experiment shows that when X is knocked out, 20 other s are not expressed How can one have such drastic effects? Regulatory Proteins Gene X
1.3 Lohkorakenne muodostetaan käyttämällä a) puolipistettä b) aaltosulkeita c) BEGIN ja END lausekkeita d) sisennystä
 OULUN YLIOPISTO Tietojenkäsittelytieteiden laitos Johdatus ohjelmointiin 811122P (5 op.) 12.12.2005 Ohjelmointikieli on Java. Tentissä saa olla materiaali mukana. Tenttitulokset julkaistaan aikaisintaan
OULUN YLIOPISTO Tietojenkäsittelytieteiden laitos Johdatus ohjelmointiin 811122P (5 op.) 12.12.2005 Ohjelmointikieli on Java. Tentissä saa olla materiaali mukana. Tenttitulokset julkaistaan aikaisintaan
Use of spatial data in the new production environment and in a data warehouse
 Use of spatial data in the new production environment and in a data warehouse Nordic Forum for Geostatistics 2007 Session 3, GI infrastructure and use of spatial database Statistics Finland, Population
Use of spatial data in the new production environment and in a data warehouse Nordic Forum for Geostatistics 2007 Session 3, GI infrastructure and use of spatial database Statistics Finland, Population
TIEKE Verkottaja Service Tools for electronic data interchange utilizers. Heikki Laaksamo
 TIEKE Verkottaja Service Tools for electronic data interchange utilizers Heikki Laaksamo TIEKE Finnish Information Society Development Centre (TIEKE Tietoyhteiskunnan kehittämiskeskus ry) TIEKE is a neutral,
TIEKE Verkottaja Service Tools for electronic data interchange utilizers Heikki Laaksamo TIEKE Finnish Information Society Development Centre (TIEKE Tietoyhteiskunnan kehittämiskeskus ry) TIEKE is a neutral,
TIETEEN PÄIVÄT OULUSSA 1.-2.9.2015
 1 TIETEEN PÄIVÄT OULUSSA 1.-2.9.2015 Oulun Yliopisto / Tieteen päivät 2015 2 TIETEEN PÄIVÄT Järjestetään Oulussa osana yliopiston avajaisviikon ohjelmaa Tieteen päivät järjestetään saman konseptin mukaisesti
1 TIETEEN PÄIVÄT OULUSSA 1.-2.9.2015 Oulun Yliopisto / Tieteen päivät 2015 2 TIETEEN PÄIVÄT Järjestetään Oulussa osana yliopiston avajaisviikon ohjelmaa Tieteen päivät järjestetään saman konseptin mukaisesti
Topologies on pseudoinnite paths
 Topologies on pseudoinnite paths Andrey Kudinov Institute for Information Transmission Problems, Moscow National Research University Higher School of Economics, Moscow Moscow Institute of Physics and Technology
Topologies on pseudoinnite paths Andrey Kudinov Institute for Information Transmission Problems, Moscow National Research University Higher School of Economics, Moscow Moscow Institute of Physics and Technology
Travel Getting Around
 - Location Olen eksyksissä. Not knowing where you are Voisitko näyttää kartalta missä sen on? Asking for a specific location on a map Mistä täällä on? Asking for a specific...wc?...pankki / rahanvaihtopiste?...hotelli?...huoltoasema?...sairaala?...apteekki?...tavaratalo?...ruokakauppa?...bussipysäkki?
- Location Olen eksyksissä. Not knowing where you are Voisitko näyttää kartalta missä sen on? Asking for a specific location on a map Mistä täällä on? Asking for a specific...wc?...pankki / rahanvaihtopiste?...hotelli?...huoltoasema?...sairaala?...apteekki?...tavaratalo?...ruokakauppa?...bussipysäkki?
FinFamily PostgreSQL installation ( ) FinFamily PostgreSQL
 FinFamily PostgreSQL 1 Sisällys / Contents FinFamily PostgreSQL... 1 1. Asenna PostgreSQL tietokanta / Install PostgreSQL database... 3 1.1. PostgreSQL tietokannasta / About the PostgreSQL database...
FinFamily PostgreSQL 1 Sisällys / Contents FinFamily PostgreSQL... 1 1. Asenna PostgreSQL tietokanta / Install PostgreSQL database... 3 1.1. PostgreSQL tietokannasta / About the PostgreSQL database...
Huom. tämä kulma on yhtä suuri kuin ohjauskulman muutos. lasketaan ajoneuvon keskipisteen ympyräkaaren jänteen pituus
 AS-84.327 Paikannus- ja navigointimenetelmät Ratkaisut 2.. a) Kun kuvan ajoneuvon kumpaakin pyörää pyöritetään tasaisella nopeudella, ajoneuvon rata on ympyränkaaren segmentin muotoinen. Hitaammin kulkeva
AS-84.327 Paikannus- ja navigointimenetelmät Ratkaisut 2.. a) Kun kuvan ajoneuvon kumpaakin pyörää pyöritetään tasaisella nopeudella, ajoneuvon rata on ympyränkaaren segmentin muotoinen. Hitaammin kulkeva
You can check above like this: Start->Control Panel->Programs->find if Microsoft Lync or Microsoft Lync Attendeed is listed
 Online Meeting Guest Online Meeting for Guest Participant Lync Attendee Installation Online kokous vierailevalle osallistujalle Lync Attendee Asennus www.ruukki.com Overview Before you can join to Ruukki
Online Meeting Guest Online Meeting for Guest Participant Lync Attendee Installation Online kokous vierailevalle osallistujalle Lync Attendee Asennus www.ruukki.com Overview Before you can join to Ruukki
KONEISTUSKOKOONPANON TEKEMINEN NX10-YMPÄRISTÖSSÄ
 KONEISTUSKOKOONPANON TEKEMINEN NX10-YMPÄRISTÖSSÄ https://community.plm.automation.siemens.com/t5/tech-tips- Knowledge-Base-NX/How-to-simulate-any-G-code-file-in-NX- CAM/ta-p/3340 Koneistusympäristön määrittely
KONEISTUSKOKOONPANON TEKEMINEN NX10-YMPÄRISTÖSSÄ https://community.plm.automation.siemens.com/t5/tech-tips- Knowledge-Base-NX/How-to-simulate-any-G-code-file-in-NX- CAM/ta-p/3340 Koneistusympäristön määrittely
Small Number Counts to 100. Story transcript: English and Blackfoot
 Small Number Counts to 100. Story transcript: English and Blackfoot Small Number is a 5 year-old boy who gets into a lot of mischief. He lives with his Grandma and Grandpa, who patiently put up with his
Small Number Counts to 100. Story transcript: English and Blackfoot Small Number is a 5 year-old boy who gets into a lot of mischief. He lives with his Grandma and Grandpa, who patiently put up with his
Kvanttilaskenta - 1. tehtävät
 Kvanttilaskenta -. tehtävät Johannes Verwijnen January 9, 0 edx-tehtävät Vastauksissa on käytetty edx-kurssin materiaalia.. Problem False, sillä 0 0. Problem False, sillä 0 0 0 0. Problem A quantum state
Kvanttilaskenta -. tehtävät Johannes Verwijnen January 9, 0 edx-tehtävät Vastauksissa on käytetty edx-kurssin materiaalia.. Problem False, sillä 0 0. Problem False, sillä 0 0 0 0. Problem A quantum state
Telecommunication Software
 Telecommunication Software Final exam 21.11.2006 COMPUTER ENGINEERING LABORATORY 521265A Vastaukset englanniksi tai suomeksi. / Answers in English or in Finnish. 1. (a) Määrittele sovellusviesti, PersonnelRecord,
Telecommunication Software Final exam 21.11.2006 COMPUTER ENGINEERING LABORATORY 521265A Vastaukset englanniksi tai suomeksi. / Answers in English or in Finnish. 1. (a) Määrittele sovellusviesti, PersonnelRecord,
6.095/ Computational Biology: Genomes, Networks, Evolution. Sequence Alignment and Dynamic Programming
 6.095/6.895 - Computational Biology: Genomes, Networks, Evolution Sequence lignment and Dynamic Programming Tue Sept 13, 2005 Challenges in Computational Biology 4 Genome ssembly 5 Regulatory motif discovery
6.095/6.895 - Computational Biology: Genomes, Networks, Evolution Sequence lignment and Dynamic Programming Tue Sept 13, 2005 Challenges in Computational Biology 4 Genome ssembly 5 Regulatory motif discovery
National Building Code of Finland, Part D1, Building Water Supply and Sewerage Systems, Regulations and guidelines 2007
 National Building Code of Finland, Part D1, Building Water Supply and Sewerage Systems, Regulations and guidelines 2007 Chapter 2.4 Jukka Räisä 1 WATER PIPES PLACEMENT 2.4.1 Regulation Water pipe and its
National Building Code of Finland, Part D1, Building Water Supply and Sewerage Systems, Regulations and guidelines 2007 Chapter 2.4 Jukka Räisä 1 WATER PIPES PLACEMENT 2.4.1 Regulation Water pipe and its
T Statistical Natural Language Processing Answers 6 Collocations Version 1.0
 T-61.5020 Statistical Natural Language Processing Answers 6 Collocations Version 1.0 1. Let s start by calculating the results for pair valkoinen, talo manually: Frequency: Bigrams valkoinen, talo occurred
T-61.5020 Statistical Natural Language Processing Answers 6 Collocations Version 1.0 1. Let s start by calculating the results for pair valkoinen, talo manually: Frequency: Bigrams valkoinen, talo occurred
FinFamily Installation and importing data (11.1.2016) FinFamily Asennus / Installation
 FinFamily Asennus / Installation 1 Sisällys / Contents FinFamily Asennus / Installation... 1 1. Asennus ja tietojen tuonti / Installation and importing data... 4 1.1. Asenna Java / Install Java... 4 1.2.
FinFamily Asennus / Installation 1 Sisällys / Contents FinFamily Asennus / Installation... 1 1. Asennus ja tietojen tuonti / Installation and importing data... 4 1.1. Asenna Java / Install Java... 4 1.2.
toukokuu 2011: Lukion kokeiden kehittämistyöryhmien suunnittelukokous
 Tuula Sutela toukokuu 2011: Lukion kokeiden kehittämistyöryhmien suunnittelukokous äidinkieli ja kirjallisuus, modersmål och litteratur, kemia, maantiede, matematiikka, englanti käsikirjoitukset vuoden
Tuula Sutela toukokuu 2011: Lukion kokeiden kehittämistyöryhmien suunnittelukokous äidinkieli ja kirjallisuus, modersmål och litteratur, kemia, maantiede, matematiikka, englanti käsikirjoitukset vuoden
HARJOITUS- PAKETTI A
 Logistiikka A35A00310 Tuotantotalouden perusteet HARJOITUS- PAKETTI A (6 pistettä) TUTA 19 Luento 3.Ennustaminen County General 1 piste The number of heart surgeries performed at County General Hospital
Logistiikka A35A00310 Tuotantotalouden perusteet HARJOITUS- PAKETTI A (6 pistettä) TUTA 19 Luento 3.Ennustaminen County General 1 piste The number of heart surgeries performed at County General Hospital
Methods S1. Sequences relevant to the constructed strains, Related to Figures 1-6.
 Methods S1. Sequences relevant to the constructed strains, Related to Figures 1-6. A. Promoter Sequences Gal4 binding sites are highlighted in the color referenced in Figure 1A when possible. Site 1: red,
Methods S1. Sequences relevant to the constructed strains, Related to Figures 1-6. A. Promoter Sequences Gal4 binding sites are highlighted in the color referenced in Figure 1A when possible. Site 1: red,
Basic Flute Technique
 Herbert Lindholm Basic Flute Technique Peruskuviot huilulle op. 26 Helin & Sons, Helsinki Basic Flute Technique Foreword This book has the same goal as a teacher should have; to make himself unnecessary.
Herbert Lindholm Basic Flute Technique Peruskuviot huilulle op. 26 Helin & Sons, Helsinki Basic Flute Technique Foreword This book has the same goal as a teacher should have; to make himself unnecessary.
Miksi Suomi on Suomi (Finnish Edition)
 Miksi Suomi on Suomi (Finnish Edition) Tommi Uschanov Click here if your download doesn"t start automatically Miksi Suomi on Suomi (Finnish Edition) Tommi Uschanov Miksi Suomi on Suomi (Finnish Edition)
Miksi Suomi on Suomi (Finnish Edition) Tommi Uschanov Click here if your download doesn"t start automatically Miksi Suomi on Suomi (Finnish Edition) Tommi Uschanov Miksi Suomi on Suomi (Finnish Edition)
Hankkeiden vaikuttavuus: Työkaluja hankesuunnittelun tueksi
 Ideasta projektiksi - kumppanuushankkeen suunnittelun lähtökohdat Hankkeiden vaikuttavuus: Työkaluja hankesuunnittelun tueksi Erasmus+ -ohjelman hakuneuvonta ammatillisen koulutuksen kumppanuushanketta
Ideasta projektiksi - kumppanuushankkeen suunnittelun lähtökohdat Hankkeiden vaikuttavuus: Työkaluja hankesuunnittelun tueksi Erasmus+ -ohjelman hakuneuvonta ammatillisen koulutuksen kumppanuushanketta
Operatioanalyysi 2011, Harjoitus 2, viikko 38
 Operatioanalyysi 2011, Harjoitus 2, viikko 38 H2t1, Exercise 1.1. H2t2, Exercise 1.2. H2t3, Exercise 2.3. H2t4, Exercise 2.4. H2t5, Exercise 2.5. (Exercise 1.1.) 1 1.1. Model the following problem mathematically:
Operatioanalyysi 2011, Harjoitus 2, viikko 38 H2t1, Exercise 1.1. H2t2, Exercise 1.2. H2t3, Exercise 2.3. H2t4, Exercise 2.4. H2t5, Exercise 2.5. (Exercise 1.1.) 1 1.1. Model the following problem mathematically:
Ihminen ja teknologia vuorovaikutuksessa. Raija Hämäläinen, JYU Kasvatustieteiden ja psykologian tiedekunta
 Ihminen ja teknologia vuorovaikutuksessa Raija Hämäläinen, JYU Kasvatustieteiden ja psykologian tiedekunta Teknologia vuorovaikutus: oppijat, käyttötarkoitus, tilat, paikka, aika.. Miten teknologian avulla
Ihminen ja teknologia vuorovaikutuksessa Raija Hämäläinen, JYU Kasvatustieteiden ja psykologian tiedekunta Teknologia vuorovaikutus: oppijat, käyttötarkoitus, tilat, paikka, aika.. Miten teknologian avulla
Kvanttilaskenta - 2. tehtävät
 Kvanttilaskenta -. tehtävät Johannes Verwijnen January 8, 05 edx-tehtävät Vastauksissa on käytetty edx-kurssin materiaalia.. Problem The inner product of + and is. Edelleen false, kts. viikon tehtävä 6..
Kvanttilaskenta -. tehtävät Johannes Verwijnen January 8, 05 edx-tehtävät Vastauksissa on käytetty edx-kurssin materiaalia.. Problem The inner product of + and is. Edelleen false, kts. viikon tehtävä 6..
Integration of Finnish web services in WebLicht Presentation in Freudenstadt 2010-10-16 by Jussi Piitulainen
 Integration of Finnish web services in WebLicht Presentation in Freudenstadt 2010-10-16 by Jussi Piitulainen Who we are FIN-CLARIN University of Helsinki The Language Bank of Finland CSC - The Center for
Integration of Finnish web services in WebLicht Presentation in Freudenstadt 2010-10-16 by Jussi Piitulainen Who we are FIN-CLARIN University of Helsinki The Language Bank of Finland CSC - The Center for
NAO- ja ENO-osaamisohjelmien loppuunsaattaminen ajatuksia ja visioita
 NAO- ja ENO-osaamisohjelmien loppuunsaattaminen ajatuksia ja visioita NAO-ENO työseminaari VI Tampere 3.-4.6.2015 Projektisuunnittelija Erno Hyvönen erno.hyvonen@minedu.fi Aikuiskoulutuksen paradigman
NAO- ja ENO-osaamisohjelmien loppuunsaattaminen ajatuksia ja visioita NAO-ENO työseminaari VI Tampere 3.-4.6.2015 Projektisuunnittelija Erno Hyvönen erno.hyvonen@minedu.fi Aikuiskoulutuksen paradigman
Returns to Scale Chapters
 Return to Scale Chapter 5.1-5.4 Saara Tuurala 26.9.2007 Index Introduction Baic Formulation of Retur to Scale Geometric Portrayal in DEA BCC Return to Scale CCR Return to Scale Summary Home Aignment Introduction
Return to Scale Chapter 5.1-5.4 Saara Tuurala 26.9.2007 Index Introduction Baic Formulation of Retur to Scale Geometric Portrayal in DEA BCC Return to Scale CCR Return to Scale Summary Home Aignment Introduction
Chapter 9 Motif finding. Chaochun Wei Spring 2019
 1896 1920 1987 2006 Chapter 9 Motif finding Chaochun Wei Spring 2019 Contents 1. Reading materials 2. Sequence structure modeling Motif finding Regulatory module finding 2 Reading materials Tompa et al
1896 1920 1987 2006 Chapter 9 Motif finding Chaochun Wei Spring 2019 Contents 1. Reading materials 2. Sequence structure modeling Motif finding Regulatory module finding 2 Reading materials Tompa et al
ELEMET- MOCASTRO. Effect of grain size on A 3 temperatures in C-Mn and low alloyed steels - Gleeble tests and predictions. Period
 1 ELEMET- MOCASTRO Effect of grain size on A 3 temperatures in C-Mn and low alloyed steels - Gleeble tests and predictions Period 20.02-25.05.2012 Diaarinumero Rahoituspäätöksen numero 1114/31/2010 502/10
1 ELEMET- MOCASTRO Effect of grain size on A 3 temperatures in C-Mn and low alloyed steels - Gleeble tests and predictions Period 20.02-25.05.2012 Diaarinumero Rahoituspäätöksen numero 1114/31/2010 502/10
make and make and make ThinkMath 2017
 Adding quantities Lukumäärienup yhdistäminen. Laske yhteensä?. Countkuinka howmonta manypalloja ballson there are altogether. and ja make and make and ja make on and ja make ThinkMath 7 on ja on on Vaihdannaisuus
Adding quantities Lukumäärienup yhdistäminen. Laske yhteensä?. Countkuinka howmonta manypalloja ballson there are altogether. and ja make and make and ja make on and ja make ThinkMath 7 on ja on on Vaihdannaisuus
Paikkatiedon semanttinen mallinnus, integrointi ja julkaiseminen Case Suomalainen ajallinen paikkaontologia SAPO
 Paikkatiedon semanttinen mallinnus, integrointi ja julkaiseminen Case Suomalainen ajallinen paikkaontologia SAPO Tomi Kauppinen, Eero Hyvönen, Jari Väätäinen Semantic Computing Research Group (SeCo) http://www.seco.tkk.fi/
Paikkatiedon semanttinen mallinnus, integrointi ja julkaiseminen Case Suomalainen ajallinen paikkaontologia SAPO Tomi Kauppinen, Eero Hyvönen, Jari Väätäinen Semantic Computing Research Group (SeCo) http://www.seco.tkk.fi/
Strategiset kyvykkyydet kilpailukyvyn mahdollistajana Autokaupassa Paula Kilpinen, KTT, Tutkija, Aalto Biz Head of Solutions and Impact, Aalto EE
 Strategiset kyvykkyydet kilpailukyvyn mahdollistajana Autokaupassa Paula Kilpinen, KTT, Tutkija, Aalto Biz Head of Solutions and Impact, Aalto EE November 7, 2014 Paula Kilpinen 1 7.11.2014 Aalto University
Strategiset kyvykkyydet kilpailukyvyn mahdollistajana Autokaupassa Paula Kilpinen, KTT, Tutkija, Aalto Biz Head of Solutions and Impact, Aalto EE November 7, 2014 Paula Kilpinen 1 7.11.2014 Aalto University
Characterization of clay using x-ray and neutron scattering at the University of Helsinki and ILL
 Characterization of clay using x-ray and neutron scattering at the University of Helsinki and ILL Ville Liljeström, Micha Matusewicz, Kari Pirkkalainen, Jussi-Petteri Suuronen and Ritva Serimaa 13.3.2012
Characterization of clay using x-ray and neutron scattering at the University of Helsinki and ILL Ville Liljeström, Micha Matusewicz, Kari Pirkkalainen, Jussi-Petteri Suuronen and Ritva Serimaa 13.3.2012
Kysymys 5 Compared to the workload, the number of credits awarded was (1 credits equals 27 working hours): (4)
 Tilasto T1106120-s2012palaute Kyselyn T1106120+T1106120-s2012palaute yhteenveto: vastauksia (4) Kysymys 1 Degree programme: (4) TIK: TIK 1 25% ************** INF: INF 0 0% EST: EST 0 0% TLT: TLT 0 0% BIO:
Tilasto T1106120-s2012palaute Kyselyn T1106120+T1106120-s2012palaute yhteenveto: vastauksia (4) Kysymys 1 Degree programme: (4) TIK: TIK 1 25% ************** INF: INF 0 0% EST: EST 0 0% TLT: TLT 0 0% BIO:
KMTK lentoestetyöpaja - Osa 2
 KMTK lentoestetyöpaja - Osa 2 Veijo Pätynen 18.10.2016 Pasila YHTEISTYÖSSÄ: Ilmailun paikkatiedon hallintamalli Ilmailun paikkatiedon hallintamalli (v0.9 4.3.2016) 4.4 Maanmittauslaitoksen rooli ja vastuut...
KMTK lentoestetyöpaja - Osa 2 Veijo Pätynen 18.10.2016 Pasila YHTEISTYÖSSÄ: Ilmailun paikkatiedon hallintamalli Ilmailun paikkatiedon hallintamalli (v0.9 4.3.2016) 4.4 Maanmittauslaitoksen rooli ja vastuut...
MRI-sovellukset. Ryhmän 6 LH:t (8.22 & 9.25)
 MRI-sovellukset Ryhmän 6 LH:t (8.22 & 9.25) Ex. 8.22 Ex. 8.22 a) What kind of image artifact is present in image (b) Answer: The artifact in the image is aliasing artifact (phase aliasing) b) How did Joe
MRI-sovellukset Ryhmän 6 LH:t (8.22 & 9.25) Ex. 8.22 Ex. 8.22 a) What kind of image artifact is present in image (b) Answer: The artifact in the image is aliasing artifact (phase aliasing) b) How did Joe
Use of Stochastic Compromise Programming to develop forest management alternatives for ecosystem services
 Use of Stochastic Compromise Programming to develop forest management alternatives for ecosystem services Kyle Eyvindson 24.3.2014 Forest Science Department / Kyle Eyvindson 3/26/2014 1 Overview Introduction
Use of Stochastic Compromise Programming to develop forest management alternatives for ecosystem services Kyle Eyvindson 24.3.2014 Forest Science Department / Kyle Eyvindson 3/26/2014 1 Overview Introduction
RINNAKKAINEN OHJELMOINTI A,
 RINNAKKAINEN OHJELMOINTI 815301A, 18.6.2005 1. Vastaa lyhyesti (2p kustakin): a) Mitkä ovat rinnakkaisen ohjelman oikeellisuuskriteerit? b) Mitä tarkoittaa laiska säikeen luominen? c) Mitä ovat kohtaaminen
RINNAKKAINEN OHJELMOINTI 815301A, 18.6.2005 1. Vastaa lyhyesti (2p kustakin): a) Mitkä ovat rinnakkaisen ohjelman oikeellisuuskriteerit? b) Mitä tarkoittaa laiska säikeen luominen? c) Mitä ovat kohtaaminen
CS284A Representations & Algorithms for Molecular Biology. Xiaohui S. Xie University of California, Irvine
 CS284A Representations & Algorithms for Molecular Biology Xiaohui S. Xie University of California, Irvine Today s Goals Course information Challenges in computational biology Introduction to molecular
CS284A Representations & Algorithms for Molecular Biology Xiaohui S. Xie University of California, Irvine Today s Goals Course information Challenges in computational biology Introduction to molecular
AYYE 9/ HOUSING POLICY
 AYYE 9/12 2.10.2012 HOUSING POLICY Mission for AYY Housing? What do we want to achieve by renting apartments? 1) How many apartments do we need? 2) What kind of apartments do we need? 3) To whom do we
AYYE 9/12 2.10.2012 HOUSING POLICY Mission for AYY Housing? What do we want to achieve by renting apartments? 1) How many apartments do we need? 2) What kind of apartments do we need? 3) To whom do we
